by Gina Belmonte (AOUS), Vincenzo Ciancia (ISTI-CNR), Diego Latella (ISTI-CNR) and Mieke Massink (ISTI-CNR)
Glioblastomas are among the most common malignant intracranial tumours. Neuroimaging protocols are used before and after treatment to evaluate its effect and to monitor the evolution of the disease. In clinical studies and routine treatment, magnetic resonance images (MRI) are evaluated, largely manually, and based on qualitative criteria such as the presence of hyper-intense tissue in the image. VoxLogicA is an image analysis tool, designed to perform tasks such as identifying brain tumours in 3D magneto-resonance scans. The aim is to have a system that is portable, predictable and reproducible, and requires minimal computing expertise to operate.
VoxLogicA is a new tool to perform image analysis in a declarative way, based on the latest developments in the theory of spatial logics [L1]. The tool was developed as part of a collaboration between the "Formal Methods and Tools" group of ISTI-CNR and the Department of Medical Physics of the Azienda Ospedaliera Universitaria Senese (AOUS).
VoxLogicA can be used for 2D (general-purpose) imaging, and for analysing 3D medical images, which has so far been the most promising application scenario. VoxLogicA has been designed with an emphasis on simplicity, and with a focus on producing explainable and implementation-independent results. A VoxLogicA session consists of a textual specification of image analysis, employing a combination of spatial features (distance between regions, or inter-reachability) with texture similarity, statistical, and imaging primitives.
VoxLogicA is publicly distributed, free and open source software (see link). At its heart lies a "model checker"; a very efficient computation engine for logical queries, exploiting advanced techniques, such as memoization and multithreading, to deliver top-notch performance.
VoxLogicA sessions are written using a declarative logical language, “Image Query Language” (ImgQL), inspired by the very successful "Structured Query Language" (SQL) for databases, but with strong mathematical foundations rooted in the area of spatial logics for topological (closure) spaces. When used in the context of medical imaging, this approach admits very concise, high level specifications (in the order of ten lines of text) that can delineate, with high accuracy, the contours of a glioblastoma tumour in a 3D Magneto-Resonance scan within eight seconds, on a standard laptop. In comparison, it takes an expert radiotherapist about half an hour to perform this task.
The same procedure has been applied to circa 200 cases (the well-known "Brain Tumour Segmentation (BraTS) challenge" dataset). Accuracy of the obtained results can be measured; the new procedure scores among the top-ranking methods of the BraTS challenge in 2017 - the state of the art in the field, dominated by machine-learning methods - and it is comparable to manual delineation by human experts. Figure 1 shows the results of segmentation of a tumour for one of those cases, where the top row shows the MRI image, the middle row the result of manual segmentation by independent experts, and the bottom row the result performed with VoxLogicA.
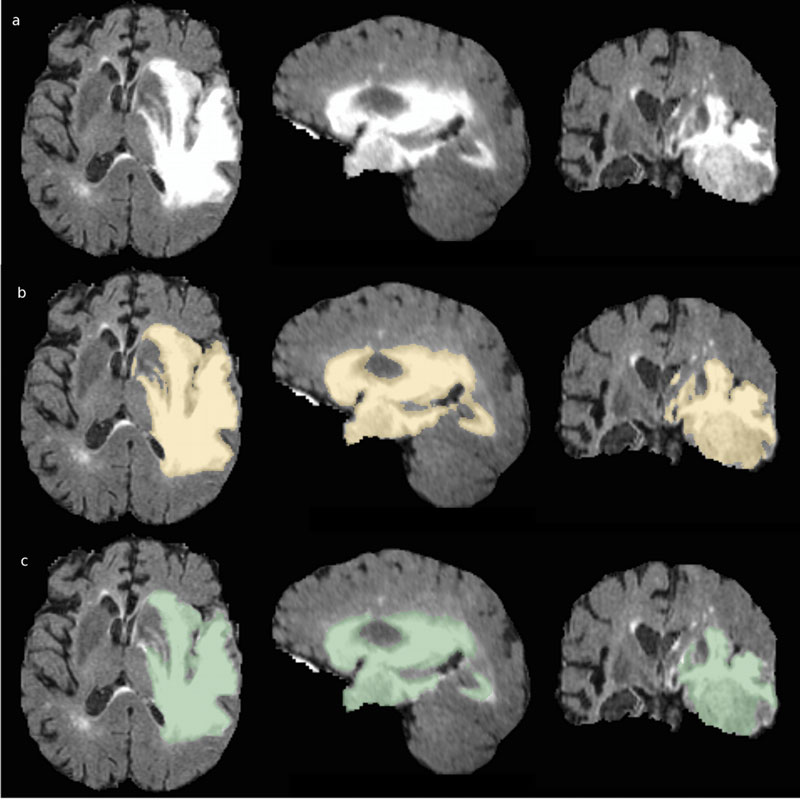
Figure 1: Results of segmentation of GTV for TCIA 471 patient: a) FLAIR acquisition b) Manual segmentation (BRATS 17 dataset) c) Segmentation result performed with VoxLogicA.
In the near future we plan to enhance this work, both in the direction of clinical case studies and to embrace other computational approaches that can be coordinated and harmonised using high-level logical specifications. Furthermore, the approach is very versatile, and its application is not limited to a single specific type of tumour or region in the body, paving the way for the analysis of other types of cancer and segmentation of various kinds of brain tissue such as white and grey matter.
A recent publication introducing the tool and its application to glioblastoma segmentation can be found in [1]. The source code and binaries of VoxLogicA are available at the link below together with a simple example of a 2D background removal task, intended as a mini tutorial for the tool. The theoretical foundations of spatial model checking can be found in [2] and an earlier study on glioblastoma segmentation performed with the general purpose spatial-temporal model checker “topochecker” can be found in [3].
Link:
[L1] https://kwz.me/hyy
References:
[1] G. Belmonte, et al.:: “VoxLogicA: a Spatial Model Checker for Declarative Image Analysis, TACAS 2019, LNCS 11427, pp. 281-298, 2019.
DOI: 10.1007/978-3-030-17462-0_16
[2] V. Ciancia, et al.:“Model Checking Spatial Logics for Closure Spaces”, Logical Methods in Computer Science Vol. 12, Nr. 4, 2016. DOI: 10. 2168/LMCS-12(4:2)2016
[3] F. Banci Buonamici, et al.: “Spatial Logics and Model Checking for Medical Imaging”, STTT, Online first, 2019. DOI: 10.1007/s10009-019-00511-9
Please contact:
Vincenzo Ciancia
CNR-ISTI, Italy











