by Sara Colantonio (ISTI-CNR), Andrea Barucci (IFAC-CNR) and Danila Germanese (ISTI-CNR)
Precision health, the future of patient care, is dependent on artificial intelligence. Of the information contained in a digital medical image, visual analysis can only extract about 10%. Radiomics aims to extract an enormous wealth of quantitative data from biomedical images, which could not be processed through simple visual analysis, but is capable of providing more information on the underlying pathophysiological phenomena and biological processes within human body. The subsequent mining of these quantitative data can offer very useful information on the aggressiveness of the disease under investigation, opening at the tailoring of the therapies based on a patient’s needs and at the monitoring of the response to care. Therefore, by using specific mathematical algorithms and artificial intelligence techniques, radiomics provides very powerful support for precision medicine, especially in oncology.
Solid tumours do not appear as a homogeneous entity, but are formed by multiclonal populations of cancer cells that exhibit considerable spatial and temporal variability. Such variability can provide valuable information on both the aggressiveness and the probability of response to therapies (therapeutic response) of the tumour itself [1].
Needle biopsy is a commonly used tool in clinical practice, but it is an invasive procedure that does not allow detecting the entire range of potential biological variation within a tumour because of the limited number of locations where cancer cells are sampled (incomplete sampling). Clinical imaging, conversely, samples the entire tumour volume in a non-invasive way and allows phenotypic characteristics to be extracted from different spatial and temporal levels, from macro-lesions up to the cellular and genetic scale.
However, image characteristics are often visually evaluated and described by radiologists or clinicians, giving rise to a subjective description of tumour imaging phenotypes, with significant intra-and inter-observer variability [2]. Moreover, only about 10 % of the information contained in a digital medical image can usually be extracted by a visual analysis.
It was within this context that radiomics was born; the answer to the search for a quantitative, objective and reproducible information extraction method for biomedical images. Underpinning radiomics [3] is computer vision, but it also has deep roots in statistics, which are often underestimated. Radiomics can be defined as the omic discipline of clinical image analysis, whose results are intrinsically objective and repeatable. An example of a radiomic pipeline is shown in Figure 1.
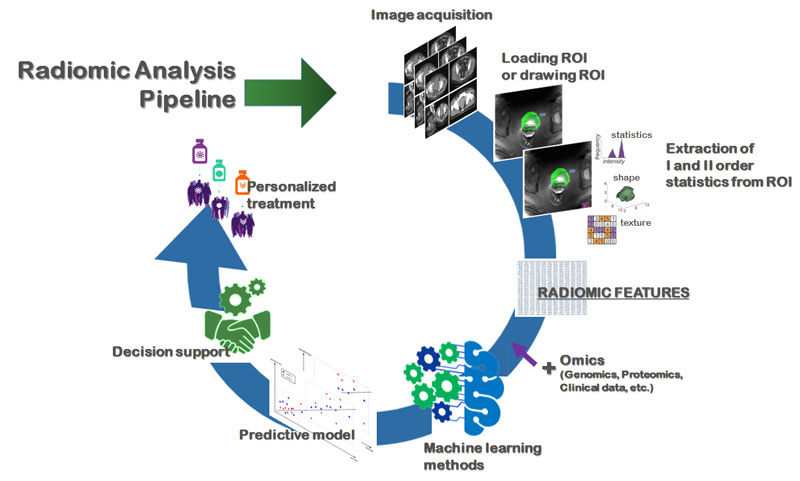
Figure 1: The pipeline of radiomic analysis toward precision medicine. For each of the imaging modalities (MRI, TC, PET, US), the first phase involves a semi-automatic or automatic segmentation of the relevant regions; this is followed by the automatic extraction of a large number of features (descriptive of the histogram, shape, texture and texture). These features are then processed with machine learning techniques, such as feature selection techniques and comparative analysis, to select the most robust ones that correlate to ground truth biomarkers (used in clinical practice).
Radiomics converts the intrinsic information within a digital image into a huge quantity of features (from a few dozen up to a few thousand) that are computed with specific mathematical algorithms. The evaluated features describe imaging parameters as intensity, shape, size, volume, textures, etc., related to the underlying tissue structures (e.g. the neoplasm and/or the surrounding healthy tissues).
By using appropriate data mining techniques (e.g. machine learning, deep learning), such features can be investigated, and the phenotypic and micro-environmental traits of a cancer tissue can be identified, enriching the data provided by laboratory tests and genomic or proteomic tests.
The identification of the “radiomic signature”, in combination with other omics data, can then be used for the development of diagnostic and prognostic models, describing phenotypic patterns connected to biological or clinical end points.
The availability of robust and validated quantitative biomarkers is fundamental to move precision medicine forward, and radiomics, in which imaging (and other) biomarkers are used to: (i) model the characteristics of each individual and the variability among individuals and (ii) predict the right treatment, for the right patient, at the right time, epitomises the research toward the implementation of personalised approaches.
Given that cancer has an intrinsically high intra- and inter-variability, owing to a range of internal and external factors, personalisation of the treatment becomes fundamental in oncology.
With this in mind, a multidisciplinary group of researchers from two CNR institutes, and clinicians from several hospitals and university centres, are combining their skills (including medical physics, biology, oncology, biomedical engineering, mathematics and computer science) to investigate the radiomic signature for the grading of prostate cancer (PCa), as well as the use of inductive representation learning methods.
Our pilot study investigated the association between radiomic features extracted from multi-parametric magnetic resonance imaging (mp-MRI), the Prostate Imaging Reporting and Data System (PI-RADS) classification, and the tumour histologic subtypes (using the pathologist Gleason score grading system), in order to identify which of the mp-MRI derived radiomic features (signature) can distinguish high and low risk PCa,, with the aim of integrating or replacing information obtained by solid biopsy.
Using a retrospective cohort of over 100 MRI patients, radiomic features (about 800) were evaluated on tumour areas segmented by the radiologists. A feed-forward method of selecting wrapper type features was used to select the four most relevant features. These were used to train (10-fold cross-validation) a narrow neural network able to predict Gleason score. Our method outperformed the majority of the works reported in literature based on standard machine learning techniques.
A descriptive paper was submitted to RSNA2020, another is undergoing submission to BIBE2019, while further data collection is underway.
References:
[1] G. Lin, K. R. Keshari, and J. Mo Park: “Cancer Metabolism and Tumor Heteroge-neity: Imaging Perspectives Using MR Imaging and Spectroscopy”, Contrast Media Mol Imaging 2017.
[2] S. Trattnig: “The Shift in Paradigm to Precision Medicine in Imaging: International Initiatives for the Promotion of Imaging Biomarkers”, Imaging Biomarkers, 2017.
[3] S. Rizzo et al.: “Radiomics: the facts and the challenges of image analysis”, European Radiology Experimental, 2018.
Please contact:
Sara Colantonio, ISTI-CNR, Italy











