by Fanny Orlhac, Charles Bouveyron and Nicholas Ayache (Université Côte d’Azur, Inria)
Radiomics is the automatic extraction of numerous quantitative features from medical images and using these features to build, for instance, predictive models. It is anticipated that Radiomics enhanced by AI techniques will play a major role in patient management in a clinical setting. To illustrate developments in this field, we briefly present two ongoing projects in oncology.
Medical images are now routinely acquired during the care pathway and play an important role in patient management. However, medical images are still largely under-exploited, using mostly visual assessment and/or the measurement of very few quantitative features available in clinical practice. To extract more information from medical images, a new field, radiomics, has successfully developed since 2010, with almost 1000 publications now using “radiomics” in PubMed, more than 30 % of which were published during the first half of 2019. Just as “genomics”, “proteomics” and “metabolomics” refer to the study of large sets of biological molecules, “radiomics” refers to the automatic extraction of large sets of quantitative features from medical images. Radiomic features can be derived from known mathematical expressions and reflect, for instance, the distribution of grey-levels, the shape of a volume of interest or the texture of the signal within that volume. More recently, in addition of these handcrafted features, an infinite number of “deep features” can be extracted from intermediate layers of convolutional neural networks.
In oncology, encouraging results using radiomic features have been published to predict biological characteristics of lesions, patient response to therapy or overall survival, for several cancer types and imaging modalities (see Figure 1). In the coming years, radiomics enhanced by artificial intelligence techniques will certainly play a major role in patient management and in the development of 4P medicine (predictive, preventive, personalised and participatory).
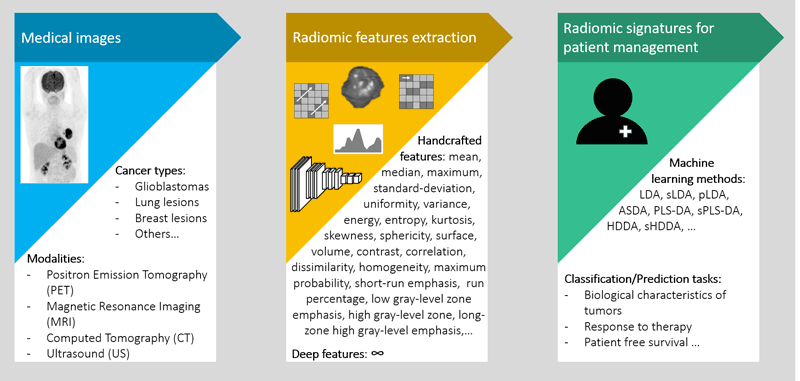
Figure 1: Process to obtain radiomic signatures in order to assist clinicians for patient management.
Epione [L1] is a research team affiliated with Université Côte d’Azur, Inria (Sophia-Antipolis, France). The long-term goal of the team is to contribute to the development of the e-patient (digital patient) for e-medicine (digital medicine). With the ongoing digital revolution in medicine and the need to analyse and interpret more and more high-dimensional data, the team has developed, for instance, a subspace discriminant analysis method which performs a class-specific variable selection through Bayesian sparsity, called sparse high-dimensional discriminant analysis (sHDDA) [1]. We demonstrated the interest of sHDDA for the radiomic analysis of computed tomography (CT) images to distinguish between lesion subtypes in lung cancer [1] or to identify triple-negative breast lesions based on positron emission tomography (PET) images [2]. Below we present two ongoing projects in radiomics in collaboration with clinical partnerships.
Management of glioblastomas
After an initial standard treatment, and in the case of clinically suspected recurrence, the differentiation between progression and radiation-induced necrosis for patients with a glioblastoma is often difficult based on magnetic resonance (MR) images. The use of PET images combined with a specific radiotracer (18F-FDOPA) improves the differential diagnosis but is still not completely accurate. In collaboration with Prof. J. Darcourt, Dr O. Humbert and Dr. F. Vandenbos from the Centre Antoine Lacassagne [L2], we studied the relevance of radiomic features extracted from PET images to distinguish between progression and radiation-induced necrosis.
Using a retrospective cohort of 78 patients with a glioblastoma, we created parametric images resulting from the subtraction of two static PET scans performed 20 and 90 minutes after injection of the radiotracer (PETsub=PET90 – PET20). Based on these new images and for each suspicious lesion, we extracted 43 radiomic features using LIFEx software [L3] including conventional features used in clinical practice, as well as histogram, shape and texture indices. We demonstrated that, thanks to a machine learning approach designed for low-sample size/high-dimensional data, the High-Dimensional Discriminant Analysis method (HDDA), it is possible to distinguish between progression and radiation necrosis with better performance than visual assessment [3]. In our cohort, the visual interpretation using the Lizarraga Scale led to a Youden Index (Y=Sensitivity+Specificity-1) of 0.27 (Sensitivity = 97 %, Specificity = 30 %). Based on the radiomic analysis of the parametric images resulting from the evolution of radiotracer uptake, the Youden Index was 0.45 (Sensitivity = 65 %, Specificity = 80 %). Additional studies are ongoing to validate these results on an independent cohort and to test if including additional features extracted from MR images could further improve the performance.
Prediction of treatment response
In oncology, a major challenge for clinicians is to identify the right treatment for the right patient at the right time. To assist them in this task, our goal is to develop radiomic signatures to predict the patient’s response to therapy. In collaboration with Dr T. Cassou Mounat, Dr A. Livartowski, and Dr M. Luporsi from Institut Curie [L4] and Dr. I. Buvat from Laboratoire IMIV [L5], we are focusing on two cancer types. In lung cancer, our objective is to combine radiomic features extracted from pre-treatment PET images in order to predict the response to chemotherapy. As nearly 50 % of patients do not respond positively, their early identification could allow clinicians to propose alternative therapies, such as immunotherapy, straight away. In breast cancer, we are developing a radiomic signature to predict the response to neoadjuvant chemotherapy. Indeed, a complete pathological response is only observed in about 20 % of patients while 10 % have stable or progressive disease after neoadjuvant chemotherapy. The early identification of non-responding patients or of tumours that will go on growing during chemotherapy would make it possible to adjust therapy at best without any loss of time.
Overall, the combination of radiomic analysis and modern machine learning approaches paves the way to better patient management thanks to a more extensive exploitation of medical images that are already currently acquired during the care pathway.
Links:
[L1] https://team.inria.fr/epione/en/
[L2] https://kwz.me/hyf
[L3] https://www.lifexsoft.org/
[L4] https://institut-curie.org/
[L5] http://www.imiv.fr
References:
[1] F. Orlhac, P.-A. Mattei, C. Bouveyron, N. Ayache: “Class-specific variable selection in high-dimensional discriminant analysis through Bayesian Sparsity”, J Chem, 2019;33:e3097.
[2] F. Orlhac, O. Humbert, T. Pourcher, L. Jing, J.-M. Guigonis, J. Darcourt, N. Ayache, C. Bouveyron: “Statistical analysis of PET radiomic features and metabolomic data: prediction of triple-negative breast cancer”, J Nucl Med, 2018;59:1755.
[3] F. Orlhac, A.-C. Rollet, C. Bouveyron, J. Darcourt, N. Ayache, O. Humbert: “Identification of a radiomic signature to distinguish recurrence from radiation-induced necrosis in treated glioblastomas using machine learning methods on dual-point 18F-FDOPA PET images”, J Nucl Med, 2019;60:57.
Please contact:
Fanny Orlhac
Université Côte d’Azur, Inria, France











