by Andrea Manno-Kovacs (MTA SZTAKI / PPKE ITK), Csaba Benedek (MTA SZTAKI) and Levente Kovács (MTA SZTAKI)
Novel 3D sensors and augmented reality-based visualisation technology are being integrated for innovative healthcare applications to improve the diagnostic process, strengthen the doctor-patient relationship and open new horizons in medical education. Our aim is to help doctors and patients explain and visualise medical status using computer vision and augmented reality.
Data from medical 3D sensors, such as computer tomography (CT) and magnetic resonance imaging (MRI), give 3D information as output, thereby creating the opportunity to model 3D objects (e.g., organs, tissues, lesions) existing inside the body. These quantitative imaging techniques play a major role in early diagnosis and make it possible to continuously monitor the patient. With the improvement of these sensors, a large amount of 3D data with high spatial resolution is acquired. Developing efficient processing methods for this diverse output is essential.
Our “Content Based Analysis of Medical Image Data” project, conducted with Pázmány Péter Catholic University, Faculty of Information Technology and Bionics (PPKE ITK) [L1], concentrated on the development of image processing algorithms for multimodal medical sensors (CT and MRI), applying content-based information, saliency models and fusing them with learning-based techniques. We developed fusion methods for efficient segmentation of medical data, by integrating the advantages of generative segmentation models, applying traditional, “handcrafted” features; and the currently preferred discriminative models using convolutional features. By fusing the two approaches, the drawbacks of the different models can be reduced, providing a robust performance on heterogeneous data, even with previously unseen data acquired by different scanners.
The fusion model [1] was introduced and evaluated for brain tumour segmentation on MRI volumes, using a novel combination of multiple MRI modalities and previously built healthy templates as a first step to highlight possible lesions. In the generative part of the proposed model, a colour- and spatial-based saliency model was applied, integrating a priori knowledge on tumours and 3D information between neighbouring scan slices. The saliency-based output is then combined with convolutional neural networks to reduce the networks’ eventual overfitting which may result in weaker predictions for unseen cases. By introducing a proof-of-concept method for the fusion of deep learning techniques with saliency-based, handcrafted feature models, the fusion approach has good abstraction skills, yet can handle diverse cases for which the net was less trained.
In a similar manner, we also implemented a technique for liver segmentation in CT scans. First, a pre-processing was introduced using a bone mask to filter the abdominal region (this is important in the case of whole-body scans). Then a combination of region growing, and active contour methods was applied for liver region segmentation. This traditional feature-based technique was fused with a convolutional neural network’s prediction mask to increase segmentation accuracy (Figure 1).
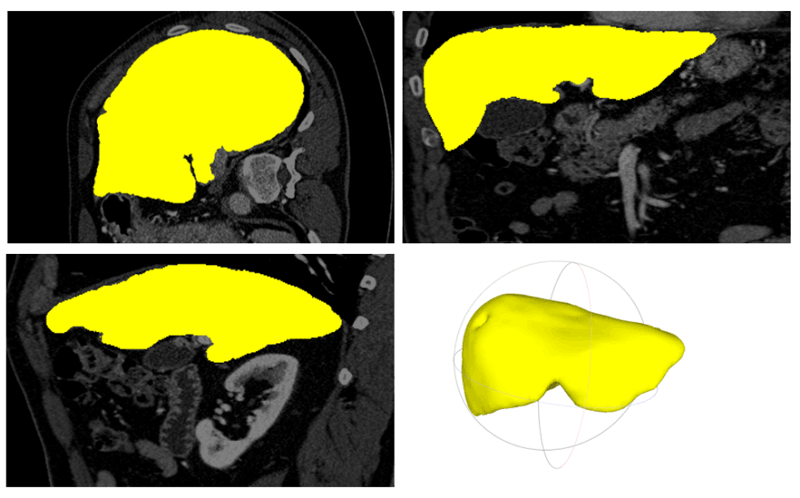
Figure 1: Liver segmentation and 3D modelling on CT data: Segmentation result in axial, coronal, sagittal view and the rendered 3D model of the liver.
The proposed techniques [2] have been successfully applied in the “zMed” project [L2], a four-year project run by Zinemath Zrt., the Machine Perception Research Laboratory in the Institute for Computer Science and Control of the Hungarian Academy of Sciences (MTA SZTAKI) [L3] together with the University of Pécs (PTE). The Department of Radiology at PTE provides input data of different modalities (CT and MRI) for the segmentation algorithms, developed by the computer vision researchers of MTA SZTAKI. Based on these tools, Zinemath Zrt is developing a software package, which provides 3D imaging and visualisation technologies for unified visualisation of medical data and various sorts of spatial measurements in an augmented reality system. By creating a completely novel visualisation mode and exceeding the current display limits, the software package applies novel technologies, such as machine learning, augmented reality and 3D image processing approaches.

Figure 2: The main motivations of the zMed project [L2].
The developed software package is planned to be adaptable to multiple medical fields: medical education and training for future physicians, introducing the latest methods more actively; improving the doctor-patient relationship by providing explanations and visualisations of the illness; surgical planning and preparation in the pre-operative phase to reduce the planning time and contributing to a more precisely designed procedure (Figure 2).
This work was supported by the ÚNKP-18-4-PPKE-132 New National Excellence Program of the Hungarian Government, Ministry of Human Capacities, and the Hungarian Government, Ministry for National Economy (NGM), under grant number GINOP-2.2.1-15-2017-00083.
Links:
[L1]: https://itk.ppke.hu/en
[L2]: http://zinemath.com/zmed/
[L3]: http://mplab.sztaki.hu
References:
[1] P. Takacs and A. Manno-Kovacs: “MRI Brain Tumor Segmentation Combining Saliency and Convolutional Network Features”, Proc. of CBMI, 2018.
[2] A. Kriston, et al.: “Segmentation of multiple organs in Computed Tomography and Magnetic Resonance Imaging measurements”, 4th International Interdisciplinary 3D Conference, 2018.
Please contact:
Andrea Manno-Kovacs
MTA SZTAKI, MPLab











