by Anna Fomitcheva Khartchenko (ETH Zurich, IBM Research – Zurich), Aditya Kashyap and Govind V. Kaigala (IBM Research – Zurich)
The role of a pathologist is critical to the cancer diagnosis workflow: they need to understand patient pathology and provide clinicians with insights through result interpretation. To do so, pathologists and their laboratory teams perform various investigations (assays) on a biopsy tissue. One of the most common tests is immunohistochemistry (IHC), which probes the expression levels for certain proteins that characterise the tissue, called biomarkers. This test enables sub-classification of the disease and is critical for the selection of a treatment modality. However, the number of biomarkers is constantly increasing, while the size of the biopsy is reducing due to early testing and more sensitive methods.
To reduce tissue consumption when performing these tests, our team implemented a variation of IHC that we termed micro-IHC (µIHC). The rationale was to reduce the area stained on the tissue section only to a region of interest at the micrometre scale, for example a small portion of the tumour. To perform µIHC we use a microfluidic probe (MFP), a liquid scanning probe, which is a device that locally deposits chemicals on specific regions of the tissue at the micrometre-length scale. This way, we perform not only local IHC staining but also can use several biomarkers on the same tissue section [1].
Using the capabilities of µIHC, we envisioned its application in optimising antibody-antigen reaction, a critical step in quality control of the IHC test. Currently, new antibodies must be tested across a range of concentrations and incubation times with multiple tissue sections to test their specificity and sensitivity. This can consume a great number of valuable samples while providing only limited information. The high tissue consumption reduces the feasibility of optimising each batch of antibodies, although it is known that they can present variations in performance. These variations are amplified when more than one tissue section is used, which often come from different sources or are prepared differently.
In these circumstances, the use of µIHC on a tissue section can generate an array with several conditions, limiting tissue consumption and equalising the whole downstream process. This potentially reduces the variability that is inherent when using different samples.
Defining “optimal” for a stain
With this methodology in hand we were faced with a rather tricky question: what is an “optimal” stain? We realised there is no straightforward answer, since the “optimal” stain varies depending on the tissue type. Take for example a common biomarker in breast cancer, HER2. The optimal stain in a healthy tissue would be “no stain”. Any stain that we observe is qualified as a false positive. However, in about 20 % of cases of breast cancer, the biomarker is present, producing a stain of varying intensities. “No stain” in this circumstance is regarded as a false negative, but even changes in stain intensity could misdirect the treatment provided by clinicians. Other staining artefacts, such as overstaining, can make the interpretation of the test more complex by masking the signal.
Using machine learning to obtain an optimal stain qualification
We asked several experts to classify our stains in “good”, “acceptable” and “not acceptable”. Nevertheless, we knew that manual labour was not an efficient and scalable way to perform this (or any) optimisation. Therefore, we decided to use the capabilities of machine learning to generate a more objective way to score the tissues, using the references provided by the pathologists [2].
We imaged all tested conditions and proceeded to extract features based on intensity, texture and the Fourier transform. These features were used for two purposes. On one side, the algorithm had to understand what kind of tissue was being considered. As mentioned, the expected staining is not the same if we are analysing a healthy tissue over a tumour tissue, making tissue identification an important facet. To do so, we train a classifier method that can learn from labelled data, a Support Vector Machine, with sets of features until we identify a set that gives the best separation between the pre-defined classes. Once this is done, the images that were not used in the training are analysed and the algorithm predicts their probability of belonging to a certain tissue type.
On the other side, the algorithm analyses the contrast level between the different compartments observed in the cell by looking at the intensities of each region. This is necessary to understand whether the stain, for the particular tissue type we are analysing, is a staining artefact. With all parts in place, the contrast level and the tissue type identifier are combined into an indicator, the “Quality Value” (QV), which gives the probability of a good quality stain ranging from 1 (best) to 0 (worst) depending on the tissue type (see Figure 1). We extracted the QVs from our analysed conditions and produced a manifold that provides essential information for optimal staining of a tissue for the patient.
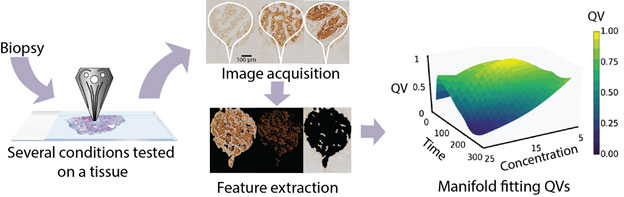
Figure 1: Schematic workflow for the processing of a tissue to obtain the quality value (QV) parameter manifold.
Outlook
The information obtained from this study can be applied on biopsy samples of individual patients to find the best staining conditions without consuming much of the sample. The remainder of the biopsy can then be used for other diagnostic tests. We believe that in this way the number of errors - defined as false positives and false negatives - caused by inadequate staining conditions may be reduced, however we are yet to decisively prove this. We hope to demonstrate such validation in future work.
We also believe the convergence of machine learning approaches with image processing and new implementations for performing biochemical assays on tissue sections will together lead to more accurate tumour profiling and thereby a more reliable diagnosis.
References:
[1] R. D. Lovchik, et al.: “Micro-immunohistochemistry using a microfluidic probe. Lab Chip 12”, 1040-1043 (2012).
[2] N. M. Arar, et al.: “High-quality immunohistochemical stains through computational assay parameter optimization”, IEEE Trans. Biomed. Eng. pp, 1–1 (2019).
Please contact:
Govind V. Kaigala
IBM Research – Zurich, Switzerland











