by Alessia Bardi (CNR-ISTI, OpenAIRE AMKE), Konstantina Galouni (ARC) and Paolo Manghi (CNR-ISTI, OpenAIRE AMKE)
The EOSC Beyond project is transforming the European Open Science Cloud into a dynamic, programmable environment by evolving the Interoperability Framework Registry from human-readable documents into machine-actionable templates. This shift empowers the scientific community to programmatically compose resources into automated scientific workflows.
The European Open Science Cloud (EOSC) is a federation of thematic and national research infrastructures with the ambition to develop a web of FAIR data and services for data-driven, Open Science. Despite the availability of a large volume of research resources in the federation, the composition of these diverse resources into integrated scientific workflows has historically remained a human-resource intensive process. To address this, the EOSC Beyond project [L1] is advancing the EOSC to support machine-composability, allowing tools and services to combine resources programmatically [1]. Central to this shift is the evolution of the EOSC Interoperability Guidelines (IF) Registry. An Interoperability Guideline (IG) is defined as a set of instructions that EOSC Providers must implement to enable specific functionalities and ensure their services or research products can work together seamlessly within the EOSC ecosystem. For example, there are interoperability guidelines for onboarding research products (metadata exchange) and for compliance with the EOSC Monitoring Service.
From Human-Readable Guidelines to Machine-Actionable Templates
In the current state of the art, the EOSC Interoperability Framework (IF) Registry primarily supports the registration of human-readable guidelines. While these were essential for human developers, they lacked the structure required for automated deployment or systemic orchestration.
The next generation of the IF Registry evolves into a collection of guidelines that support the programmatic composition of resources. This is achieved through two new technical pillars:
- Configuration Templates: These are structured metadata profiles that allow providers of guidelines to detail which information must be provided by a resource (e.g. a service) that is compliant with the guideline. For example, the IG for onboarding research products require the service to specify the base URL of its OAI-PMH endpoint and other details required for the harvesting.
- Configuration Instances: These are the specific, populated parameters provided by a service that indicate how it complies with a given template. For example, if the provider registers a service and declares it is compliant with the IG for onboarding research products, then it must provide the actual OAI-PMH endpoint, and the other required details, such as the metadata format it supports.
By recording these machine-actionable descriptions in the EOSC Service Registry, the framework enables core components and third-party applications to interact with services without manual intervention.
Case Study: The Scholarly Communication Metadata Aggregator
The power of this machine-actionable approach is best demonstrated by the Scholarly Communication Metadata Aggregator for EOSC Nodes [L2]. This application highlights how machine-readable guidelines can automate scholarly communication workflows that previously required human intervention.
In a traditional setup, a user would need to manually find a repository, identify its OAI-PMH endpoint, and configure a harvesting tool. Under the new framework, the process is fully automated:
- Discovery: The application queries the Service Registry to identify data sources that are compliant with the Interoperability Guidelines for onboarding research products (see Figure 1).
- Automated Configuration: Instead of manual setup, the application reads the Configuration Instance of the selected service, which contains the exact parameters needed to harvest its metadata.
- Execution: The application launches the harvesting process automatically based on these machine-read parameters.
- Delivery: Once the process is complete, the user is automatically notified via email with a link to download the aggregated metadata records.
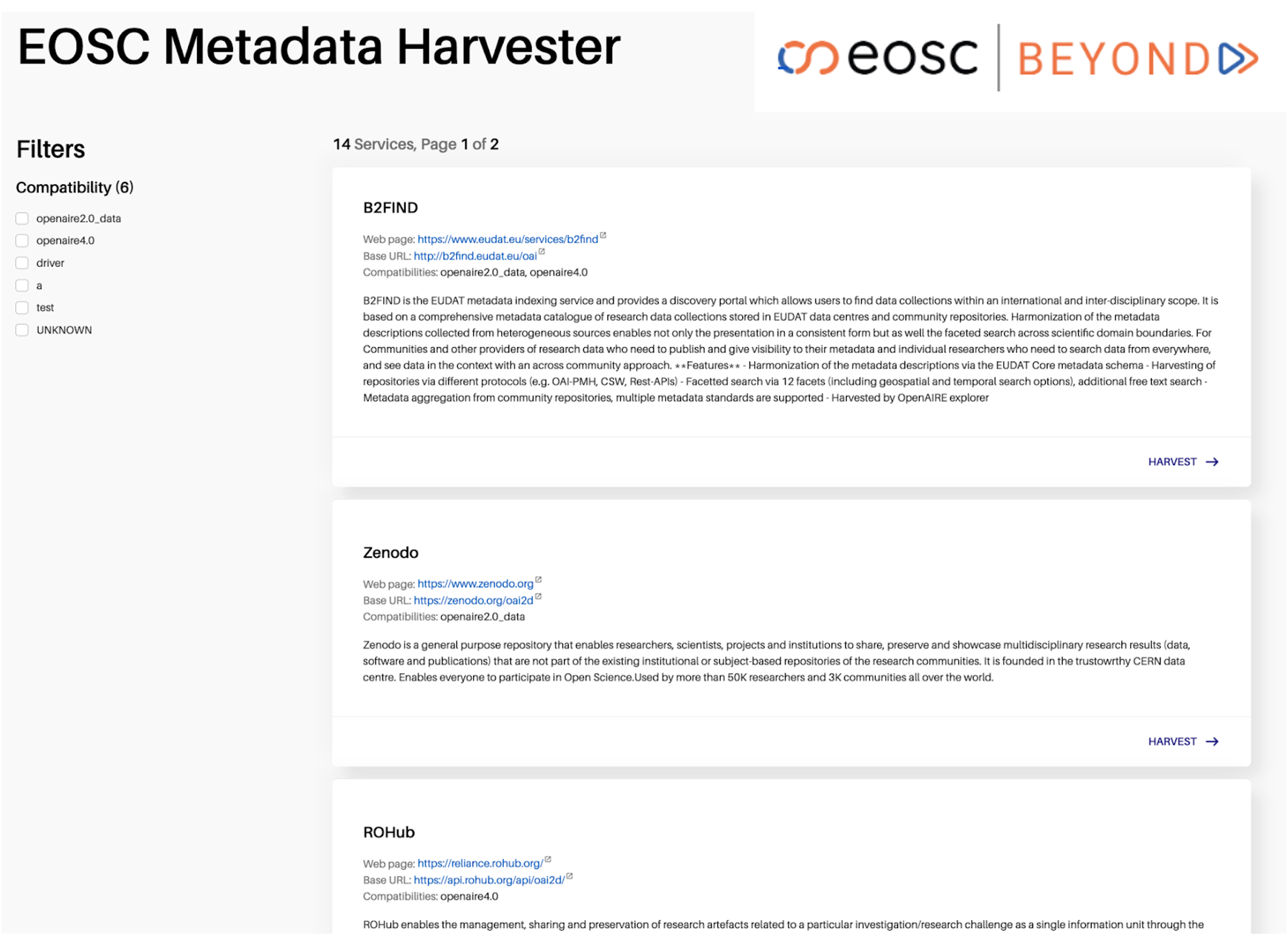
Figure 1: The Scholarly Communication Metadata Aggregator demo application.
Conclusion and Future Impact
EOSC Beyond is creating a "Web of FAIR Data and Services" that is composable in automated workflows. The framework reduces the technical burden on researchers and service providers alike, allowing them to focus on research rather than integration code. The Scholarly Communication Metadata Aggregator demonstrates the potential of machine-actionable interoperability guidelines for the realisation of automated or semi-automated workflows. The EOSC Beyond technical roadmap includes further enhancements both to the framework and the use case application to improve the user experience and integrate other services of the EOSC ecosystem, such as the EGI Data Transfer for safe transfer of harvested records in a remote location selected by the user.
Acknowledgment: This work is partly founded by the European Union EOSC Beyond project grant agreement 101131875. Views and opinions expressed are however those of the author(s) only and do not necessarily reflect those of the European Union. Neither the European Union nor the granting authority can be held responsible for them.
Links:
[L1] https://www.eosc-beyond.eu/
[L2] https://code-repo.d4science.org/MaDgIK/eosc-metadata-harvester
Reference:
[1] Bardi, A., Brandt, C., Caballer, M., Moltó, G., & Mantes, T. (2025). EOSC Beyond D13.2 First report on EOSC Execution Framework Architecture and capabilities. Zenodo. https://doi.org/10.5281/zenodo.17234674
Please contact:
Alessia Bardi, CNR-ISTI, OpenAIRE, Italy










