by Adrián Segura Ortiz, José García-Nieto (ITIS Software, University of Málaga) and Laetitia Jourdan (CRIStAL Laboratory, University of Lille)
MOEBA-BIO is an open and extensible evolutionary biclustering framework that empowers the biomedical research community to specialize in advanced optimisation strategies. Through its modular design, holistic representation of solutions, adaptive learning of the number of biclusters, and integrated self-configuration mechanisms, the framework enables expert-driven domain refinement while promoting sustainable open research software.
From Algorithm to Open Research Software
Open Science increasingly requires that computational methods are not only published, but shared as reusable, extensible and sustainable research artefacts. In computational biomedicine, however, advanced optimisation techniques are often released as standalone algorithms tightly coupled to specific studies. While scientifically valuable, such implementations can be difficult to extend, adapt to new domains, or integrate into broader research infrastructures.
Transforming algorithmic contributions into open research software requires architectural decisions that prioritise modularity, interoperability and long-term maintainability from the outset. In this context, researchers at the University of Málaga (Spain), within the Khaos Research Group [L1] and the ITIS Software Institute [L2], developed MOEBA-BIO [1] in close collaboration with the ORKAD research team [L3] of the CRIStAL laboratory [L4], University of Lille (France), as an open framework for evolutionary biclustering in biomedical data analysis. Rather than delivering a single method, it provides a configurable infrastructure in which representations, objective functions, evolutionary operators and optimisation strategies are separated into independent components. This modular architecture allows researchers to assemble, extend and refine evolutionary biclustering strategies without redesigning the entire system. The overall architecture of the framework and its modular objective space are illustrated in Figure 1.
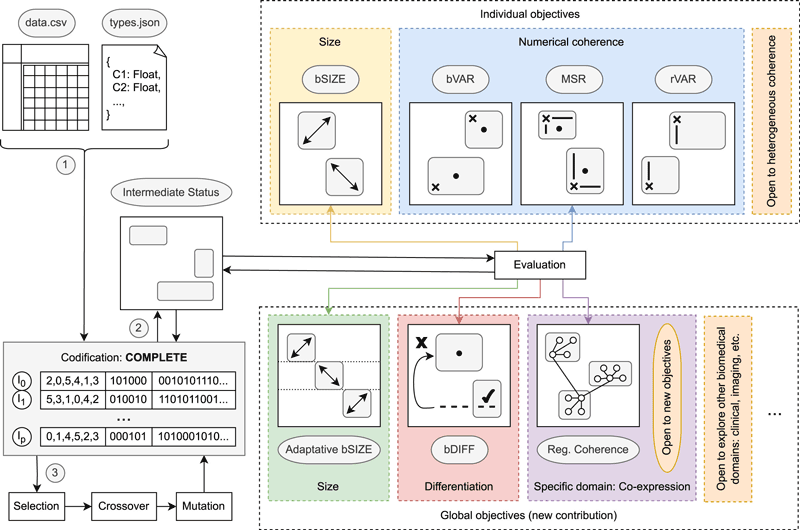
Figure 1: Overview of the MOEBA-BIO framework architecture. The workflow begins with structured biomedical input data and a column-type specification, which are transformed into a holistic solution representation (“COMPLETE” encoding). Each individual models a full biclustering structure and evolves through selection, crossover and mutation. The evaluation stage integrates traditional individual objectives (e.g., size and numerical coherence) together with newly introduced global objectives such as adaptive bicluster size, structural differentiation and domain-specific regulatory coherence. The modular design allows the integration of additional objectives and the exploration of heterogeneous biomedical domains, supporting extensibility and expert-driven specialisation within an open research software ecosystem.
A Holistic Representation for Structural Learning
A central design decision of the framework is its holistic representation of solutions. Instead of encoding individual biclusters independently, each candidate solution models a complete structure. This shift enables evaluation strategies that consider not only the internal quality of each bicluster, but also the global organisation and coherence of the solution as a whole.
This global perspective makes it possible to design objective functions that evaluate solution-wide properties, such as differentiation between groups, overall coverage or coherence with biological constraints. Rather than optimising isolated substructures, the framework supports modelling decisions that reflect the systemic nature of biomedical data. Evolutionary search thus becomes structurally aware instead of purely local.
Adaptive Learning of the Number of Biclusters
One direct consequence of this holistic representation is the adaptive determination of the number of biclusters. Traditional biclustering workflows often require this parameter to be fixed in advance or selected through external heuristics [2]. In MOEBA-BIO, however, the quantity of biclusters evolves dynamically as part of the optimisation process.
By embedding solution size into the evolutionary encoding, model complexity becomes an optimisable dimension. The algorithm can determine how many coherent groups best describe the data under the selected objectives, without imposing artificial constraints. This is particularly relevant in biomedical applications, where predefined structural assumptions may bias interpretation or overlook latent biological organisation.
This capability reduces the need for post-processing or manual adjustment and promotes methodological transparency. Because the number of groups is learned under the same optimisation criteria that guide structural coherence and domain-specific objectives, the resulting models remain internally consistent.
Supervised and Unsupervised Self-Configuration
Beyond structural learning, MOEBA-BIO incorporates a dedicated self-configuration mechanism designed to facilitate domain-driven specialisation. This component operates in two complementary phases.
In a supervised configuration phase, the framework evaluates alternative combinations of objectives and algorithmic components against representative benchmark datasets for which reference structures are available. Using wrapper-based optimisation, it identifies configurations that best align optimisation behaviour with domain expectations. This allows the framework to automatically select and weight objective functions in a way that reflects the characteristics of a specific biomedical context.
Once this higher-level configuration is established, an unsupervised refinement phase further tunes optimisation parameters such as evolutionary operators and algorithm settings. This second stage focuses on improving convergence and solution quality without relying on external labels, ensuring adaptability to new datasets while preserving generality.
Together, these supervised and unsupervised layers transform the framework into an adaptive analytical environment. Domain experts can guide specialisation through representative data, while the internal optimisation process refines performance autonomously. This combination reduces technical barriers and supports expert-driven domain refinement without sacrificing reproducibility.
Open, Interoperable and Extensible by Design
MOEBA-BIO was developed following open research software principles. The framework is fully open source, documented and publicly available through GitHub [L5]. Its architecture encourages extension: new objective functions, representations or evolutionary operators can be integrated without modifying the core system.
To promote interoperability, the framework is distributed through the Maven Central Repository [L6], enabling seamless integration into Java-based research pipelines and larger analytical infrastructures. This design choice supports sustainable reuse, reproducible workflows and incorporation into educational environments.
By clearly separating representation, evaluation and optimisation layers, the framework enhances transparency and traceability. Researchers can inspect, modify and extend each component independently, strengthening methodological openness.
Demonstrated Specialisation in Gene Co-Expression
The extensibility of the framework has already been demonstrated in gene co-expression analysis. By incorporating a regulatory coherence objective derived from inferred gene regulatory networks [3], MOEBA-BIO was specialised to align biclustering results with biologically meaningful modular structures.
Importantly, this domain adaptation did not require redesigning the system. Instead, new objectives were integrated into the existing architecture, illustrating how the framework enables community-driven extension rather than isolated reimplementation.
Towards Sustainable Evolutionary Research Infrastructures
As biomedical datasets grow in complexity and heterogeneity, analytical tools must evolve beyond fixed algorithms. Open frameworks that enable structural learning, adaptive configuration and domain-driven specialisation are essential for sustainable scientific progress.
MOEBA-BIO illustrates how evolutionary optimisation can transition from isolated methodological contributions to reusable, extensible research infrastructure. By embedding modularity, interoperability and expert-guided adaptability into its core design, the framework aligns with broader Open Science priorities centred on transparency, sustainability and collaborative development.
Links:
[L1] https://khaos.uma.es/
[L2] https://itis.uma.es/
[L3] https://orkad.univ-lille.fr/
[L4] https://www.cristal.univ-lille.fr/
[L5] https://github.com/AdrianSeguraOrtiz/MOEBA-BIO
[L6] https://mvnrepository.com/artifact/io.github.adrianseguraortiz/moeba-bio
References:
[1] A. Segura-Ortiz, et al., “Exhaustive biclustering driven by self-learning evolutionary approach for biomedical data,” Computer Methods and Programs in Biomedicine, p. 108846, 2025.
[2] M. Vandromme, et al., “A biclustering method for heterogeneous and temporal medical data,”
IEEE Transactions on Knowledge and Data Engineering, vol. 34, no. 2, pp. 506–518, 2020.
[3] A. Segura-Ortiz, et al., “Multifaceted evolution focused on maximal exploitation of domain knowledge for the consensus inference of Gene Regulatory Networks,” Computers in Biology and Medicine, vol. 196, p. 110632, 2025.
Please contact:
Adrián Segura Ortiz
ITIS Software, University of Málaga, Spain
José Francisco Aldana Montes
ITIS Software, University of Málaga, Spain











