by Ercan Engin Kuruoglu (ISTI-CNR) and Yang Li (Tsinghua-Berkeley Shenzhen Institute)
Daily estimates of parameters relating to epidemics can help public health experts to track the changes in the epidemic’s dynamics and the effectiveness of public health policy changes.
It’s approaching one year since COVID-19 first emerged: a year that is rich with stories of success and failure, contributed to by various factors. The heterogeneous temporal and spatial characteristics of data relating to the epidemic reflect various factors including government health policies, cultural differences between societies, public responses to epidemic control policies, and possibly mutations that affect the transmissibility of the virus. Temporal heterogeneity is particularly notable owing to the rather drastic changes in public policy over the time – such as lockdowns and reopening period during the summer. The delay in the response of the epidemic curve to public policy changes also reflects the response of society or how strictly the policies are enforced. Research groups around the world have modelled the spread of virus using classical compartmental models such as SIR and SEIR. Most of these works unfortunately assume fixed parameters for the epidemic models. The fixed parameter models fall short of describing the non-smooth curves demonstrated by epidemic’s spread in almost every country.
The epidemic’s parameters can give public health authorities important information regarding the changes in the transmissibility of the virus and the effect of changes in control policies. In particular, the parameters obtained from each city’s local data can point to problems associated with implementing policies, such as the local lockdowns in regions in Italy. Fixed parameter models do not provide the temporal information needed to help public health officials to adjust or fine-tune policies. It is therefore vital to develop methods that can track epidemic’s parameters daily.
This challenge was addressed by researchers from ISTI-CNR, China, and Data Science and Information Technology Center of Tsinghua-Berkeley Shenzhen Institute, China. We adopted a state-space formulation that relates temporal observations to hidden parameters of the epidemic. A statistical modelling approach was used, which aims to estimate the probability distributions of hidden variables, rather than just point estimates. This formulation also makes it possible to handle non-Gaussian distributions and nonlinear dynamics in data. Although the classical solution for this problem is provided by Kalman filtering, it necessarily makes the assumption of linearity of the system and Gaussianity of state and observation noise.
We have utilised the Unscented Kalman Filter for the solution of the state-space model which both deals with non-Gaussianity and nonlinearity, and offers computational simplicity. Despite its name, the Unscented Kalman Filter (UKF) has a different computing approach to the Kalman filter. It follows a sampling approach and avoids matrix calculations. The method is also faster when compared to other sampling-based approaches, such as sequential Monte Carlo.
We have applied this method, which provides daily estimates of transmissibility rate and error bounds, to daily epidemic data from the USA, UK, Italy and Turkey. It has demonstrated precise fits to the actual data: the methods were even capable of capturing daily changes in data from Italy during the summer period, showing the patterns of holiday-making during weekends. The method can also be used to make short-term predictions for the coming days. Figure 1a shows the daily statistics for Italy and the fit to it by UKF. In Figure 1b, we provide the corresponding daily transmissibility index for Italy calculated by UKF. This shows the drastic effect that occurred within a week of lockdown after 10 March 2020 and the rise of infected cases again within one week of normalisation (the removal of travel restrictions) on 3 June 2020.
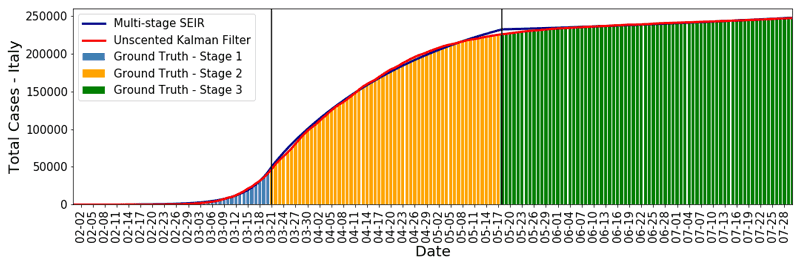
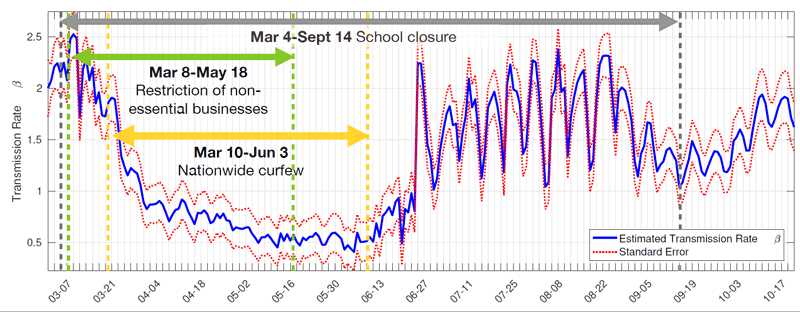
Figure 1: a) Daily epidemic statistics for Italy and the fit to it by UKF (top); b) The corresponding daily transmissibility index for Italy calculated by UKF (bottom).
As we continue this project, we will analyse the data from different regions of Italy and new factors will be introduced to the model first to take care of the period for immunity and also in anticipation of the arrival of a vaccine. We will also include the availability of healthcare resources as a parameter in our model and will simulate extreme case scenarios in which the health system has been stretched beyond capacity.
The software we have developed is available to public health officials upon request. We welcome experts on epidemics and health officials to contact us to collaborate.
Please contact:
Ercan Engin Kuruoglu, ISTI-CNR, Pisa, Italy











