by Fabrice Jouanot (Univ. Grenoble Alpes), Olivier Palombi (Univ. Grenoble Alpes, CHU de Grenoble) and Marie-Christine Rousset (Univ. Grenoble Alpes, IUF)
Through SIDES 3.0, we are developing an ontology-based e-learning platform in medicine to make learning analytics transparent and explainable to end-users (learners and teachers). In this project, the educational content, the traces of students' activities and the correction of exams are linked and related to items of an official reference program in a unified data model. As a result, an integrated access to useful information is provided for student progress monitoring and equipped with a powerful query language allowing users to express their specific needs relating to data exploration and analysis.
The goal of SIDES 3.0 is to empower medical students and teachers in data analytics to enable them to take charge of their own progress monitoring and training.
We have followed the Semantic Web [1] and Linked Data [2] standards for the semi-automatic and modular construction of OntoSIDES [3], a knowledge graph comprising a lightweight domain ontology serving as a pivot high-level vocabulary of the query interface with users, and of a huge dataset that is automatically extracted from SIDES dumps using mappings. Both the ontological statements and the factual statements are uniformly described in RDF format [L1], which makes it possible to query OntoSIDES using the SPARQL query language [L2].
It is important to note that no personal data concerning students are extracted from SIDES. Instances of students in OntoSIDES are just represented by identifiers of the form sides:etu12402 (as shown in Figure 1).
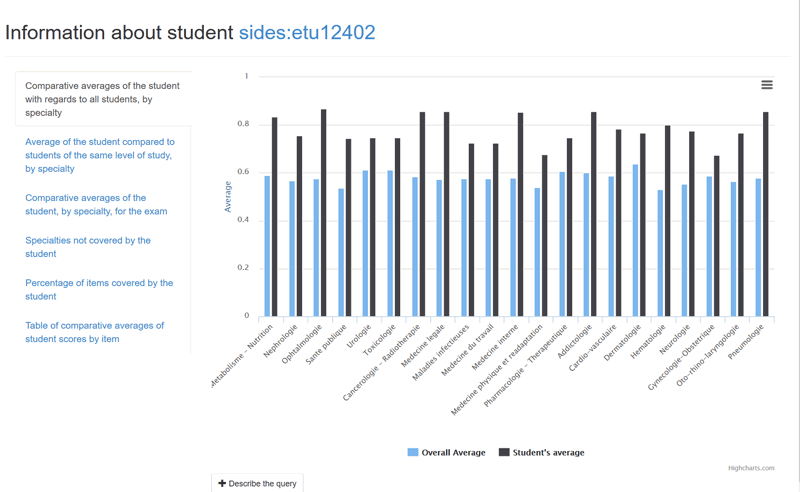
Figure 1: Parametrized queries for students.
The current version of the OntoSIDES knowledge graph has scaled up to 6,5 billion RDF triples describing training and assessment activities performed by more than 145,000 students over almost six years on the SIDES French national e-learning platform. In OntoSIDES, exams and training tests are made up of multiple choice questions and student activities are described at the granularity of time-stamped clicks of students’ selected answers to each question.
Through SPARQL queries, students can explore parts of the program that they haven’t yet studied, or launch a comparative analysis of their own progress in a specific part of the program. The same mechanisms let teachers analyse the strengths and weaknesses of their class compared to other groups at the same study level in other universities.
Since we do not expect end-users to master the raw syntax of SPARQL and to express directly complex queries in SPARQL, we have designed a set of parametrised queries that users can customise through a user-friendly interface. Figure 1 shows the interface for students who can choose to launch one of the queries in the left column that will be parametrised by the student’s identifier (sides:etud12402 in the example). The chart in the right side of the figure shows the result returned to the first query for that student, i.e., the average results obtained per specialty by the student (to all the questions he/she answered related to this specialty), compared to the overall average obtained per specialty by the whole group of active students.
This methodology is not specific to medicine ; it can be applied to other disciplines for enriching or building learning management systems. The effort required depends mainly on the existence of a reference standard for the educational program in the target discipline.
SIDES 3.0 is an ongoing project funded by the French Programme Investissement d’Avenir (PIA) through the ANR call DUNE (Développement d'Universités Numériques Expérimentales). It is conducted under the authority of UNESS (Université Numérique en Santé et en Sport) and involves Université Grenoble Alpes, Inria, and Ecole Normale Supérieure as partners. It is built upon the SIDES e-learning platform, which has been used since 2013 by all medical schools in France for online graduation and assessment.
Links:
[L1] http://www.w3.org/TR/rdf-concepts/
[L2] https://www.w3.org/TR/
References:
[1] D. Allemang, J. Hendler: “Semantic Web for the Working Ontologist: Modeling in RDF, RDFS and OWL”, Morgan Kaufmann, 2011.
[2] T. Heath, C. Bizer: “Linked Data : Evolving the Web into a Global DataSpace”, Morgan and Claypool, 2011.
[3] O. Palombi, et al.: “OntoSIDES: Ontology-based student progress monitoring on the national evaluation system of French Medical Schools”, Artificial Intelligence in Medicine, vol 96, 2019.
Please contact:
Marie-Christine Rousset
Univ. Grenoble Alpes, France











