by Georgios Stamatakos
Approaching biology as the physical science of living matter dictates the development of a parsimonious mathematical and computational formulation of multiscale biological phenomena. Such a long-term endeavour must be collaborative on a worldwide scale. The combination of cancer biology with in silico oncology can serve as a valuable paradigm for such a process. Here we outline simulation results on the response of tumorous and normal tissues to therapeutic schemes. These simulations were developed over the last decade by the In Silico Oncology Group at the National Technical University of Athens.
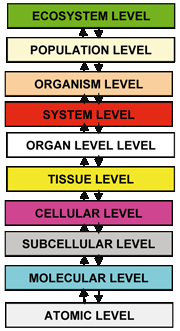
The remarkable rate of accumulation of both experimental and observational (clinical) knowledge pertaining to living matter dictates the formulation of a parsimonious system of "laws" that are somewhat analogous to Newton's "Mathematical Principles of Natural Philosophy" This seems to be a necessary step if a rational, coherent and transparent understanding of the biological phenomena is to be sought. Such a system would consist of a finite yet considerable number of principles and would refer to all levels of biocomplexity (see Figure 1), since according to Denis Noble a privileged level of causality does not appear to exist. The experimental, observational and theoretical study of cancer, a markedly multiscale biological phenomenon of obvious clinical importance, may be viewed as an excellent ground for the establishment of a number of such multiscale laws. Their formulation might well be achieved in a combination of discrete and continuous mathematical terms.
In Silico Oncology and the Problem of Cancer Predictability
The emerging field of in silico (computational) oncology has already provided some plausible descriptions of several biological mechanisms, both continuous and discrete in nature, that characterize cancer. Obviously cancer is far from being a purely deterministic phenomenon. Instead it seems to behave like a mixture of deterministic (eg sequence of cell-cycle phases) and stochastic (eg radiation cell-kill probability) processes. Stochastic aspects should therefore always be taken into account. Nevertheless, as more critical knowledge becomes available, the more deterministic the cancer phenomenon appears to become. An illustrative example supporting this hypothesis is that more detailed knowledge of the genetic status of a tumour may lead to a better prediction of its response to therapeutic interventions, and thus to apparently more deterministic tumour behaviour.
Based on these ideas, the In Silico Oncology Group (ISOG) in the Institute of Communication and Computer Systems (ICCS) at the National Technical University of Athens (NTUA) has developed a number of hybrid discrete Monte Carlo/cellular automata and continuous differential equation simulation models of tumour growth, and of the response of tumour and normal tissues to therapeutic modalities. The models range from tumour growth and radiotherapy response in vitro, to the clinical tumour response, to radiotherapeutic and chemotherapeutic schemes in vivo, based among other things on actual imaging data. Processed molecular data is used in order to perturb the radiobiological or pharmacodynamic cell-kill parameters about their population-based mean values. At the heart of the proposed simulation approach lies a prototype system of quantizing cell clusters included within each geometrical cell of a discretizing mesh, which covers the anatomic area of interest. Cell-cycle phase durations and imaging-based metabolism distribution define the quantization equivalence classes considered. Several algorithms have been developed so as to simulate various macroscopic mechanisms such as tumour expansion or shrinkage and mechanical boundary conditions, as well as the effects of particular drugs (eg temozolomide) and radiation on the tumorous and normal tissue under consideration.
A number of the models developed, which mainly refer to imageable glioblastomas, have already been clinically validated to a substantial degree by exploiting the outcomes of pertinent clinical trials. Long-term clinical testing and adaptation procedures are in progress. The response of treatment-affected normal tissues in radiotherapeutic schemes has also been addressed for certain cases. Currently, a substantial extension of the simulation models to cases of nephroblastoma (Wilm's tumour) and breast cancer is being performed within the frame of the EC-funded project ACGT (Advancing Clinico-Genomic Trials on cancer), in collaboration with several European institutions including the Foundation for Research and Technology Hellas, Heraklion, in Greece. Of particular clinical importance is the tight collaboration with the Paediatric Haematology and Oncology Clinic of the University of Saarland in Germany, and Belgium's Institut Jules Bordet, located in Brussels. The whole effort is also supported by the NIH-NCI-funded Center for the Development of a Virtual Tumor (CviT), based in Massachusetts, USA. It is worth noting the remarkably collaborative character of this and other complementary research efforts on a global scale.
It is expected that the type of model described here will provide clinicians and researchers with the option of running virtual experiments to optimize cancer treatment strategies based on the specific molecular, histopathologic, imaging and historical data of individual patients. A deeper understanding of the cancer disease at a molecular level and at the same time of the related macroscopic phenomena is a further intermediate goal of considerable importance.
Links:
In Silico Ontology group, NTUA: http://www.in-silico-oncology.iccs.ntua.gr
ACGT project: http://www.eu-acgt.org/
Center for the Development of a Virtual Tumor: https://www.cvit.org/
Please contact:
Georgios Stamatakos
National Technical University of Athens, Greece
Tel: + 30 210 772 2288
E-mail: gestam![]() central.ntua.gr
central.ntua.gr










