by Sara Colantonio, Davide Moroni and Ovidio Salvetti
Researchers in the EU HEARTFAID consortium are undertaking an investigation into the extraction and deployment of new features for the representation of cardiac shape and function.
The ISTI-CNR "Signals and Images" Laboratory is currently involved in a challenging research activity concerning cardiac image analysis. This is taking place in the context of the EU-funded project HEARTFAID. This project aims at providing a knowledge-based platform of services for supporting the clinical management of heart failure. Here we report the preliminary results of our investigation.
The analysis of deformation patterns in the heart is of key importance in understanding its functional properties and assessing its state of health. Image analysis modalities provide an invaluable aid when studying complex cardiac structures. However, image sequences contain a huge amount of high-dimensional data (two or three spatial dimensions plus time), which cannot be fully exploited without the help of suitable tools for image processing and pattern recognition. Furthermore, the growing number of imaging perspectives and modalities provide multi-source information on the anatomical structures, electromagnetic activity, dynamic perfusion and metabolism, strain and blood flow of the heart, which must somehow be combined into an overall picture. There is thus a pressing need for a unifying framework for cardiac dynamic analysis.
Cardiac modelling seems to be the natural answer. An abstract representation of the heart is built and can be instantiated to the particular anatomy under examination, with the aim of extracting shape and functional parameters. In addition, cardiac modelling can enable a sophisticated quantitative assessment of heart pathologies, for which present clinical practice uses only semi-quantitative (and to some extent subjective) measurements.
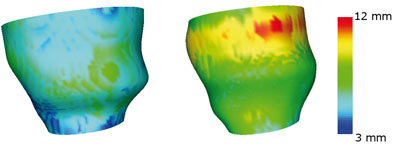
In the clinical management of heart failure patients, the study of segmental wall motion and dyssynchrony characterization are so far the main areas in which image processing has proven useful or even essential. In particular, dyssynchrony, which refers to incoordinate wall motion due to activation delay, is a complex phenomenon whose origins are to be tracked back to electrical conduction disturbances that affect both regional and global functions of the heart. Despite its relevance, the only dyssynchrony marker that has received some consensus is an ECG-derived parameter, yet this is poorly correlated to the outcome of resynchronization therapy. We believe that cardiac modelling would provide more insight into the problem by conveying novel representation features and suitable tools for their scientific visualization. Ultimately, dyssynchrony characterization may be translated via the extracted features into a statistical pattern-recognition problem, thus allowing for new methods of quantification.
With this goal in mind, we investigated approaches that describe periodically deforming structures identified in 3D image sequences (MRI, ultrafast CT etc) in a compact but faithful fashion. An encoding of this type would be useful to build up a reference database for similarity searches or data mining procedures.
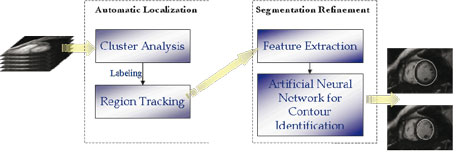
Of course an essential step in characterizing deformable structures is their initial localization and reconstruction from an image sequence. We addressed this preliminary but highly nontrivial problem by developing a two-stage procedure, based on fuzzy clustering and Artificial Neural Networks (ANNs), for the identification and reconstruction of the deformable structure of interest.
The reconstructed structures represent the underlying geometry of the organ, which is further enriched by computing local attribute functions, describing its functional properties. A model normalization procedure is then applied to solve issues of data compression, enable easy comparison of deformable structures belonging to different patients and, finally, to identify the most salient features. Both Fourier and wavelet transforms with respect to time are considered, in order to get an overall encoding of the whole deformation cycle.
Future activities will involve fine tuning of the encoding with respect to the characterization of wall motion and dyssynchrony, with the goal of identifying meaningful nontraditional descriptors. In this setting, the HEARTFAID platform could provide a unique opportunity to start their comparative evaluation.
The HEARTFAID consortium consists of four technological research institutes (University of Calabria-Italy, the coordinator; FORTH-Greece; ISTI-CNR-Italy and Rudjer Boskovic Institute-Croatia), four medical partners (University of Milano Bicocca-Italy, Istituto Auxologico-Italy, University Magna Graecia-Italy and Jagiellonian University Medical College-Poland), and three industrial partners (FORTHnet SA-Greece, Synapsis-Italy and VMWS-UK). The project began in February 2006 and will run for three years.
Links:
HEARTFAID project http://www.heartfaid.org/Signals & Images Laboratory, ISTI-CNR: http://www.isti.cnr.it/ResearchUnits/Labs/si-lab/
Please contact:
Davide Moroni
ISTI-CNR, Italy
Tel: +39 050 315 3130
E-mail: davide.moroni![]() isti.cnr.it
isti.cnr.it










