by Lakshmi Sastry and Srikanth Nagella
The Integrative Biology (IB) project, funded by EPSRC, is nearing completion with the building of a customized Grid framework. This is being used to run large multi-scale models, from cellular to whole organ simulations, to manage a growing database of in-vitro experimental data and simulation results, and to support advanced visualization for interactive data analysis with comparison and assimilation of experimental and observed data. The services offered by the computational framework are based on the requirements of two application areas: arrhythmia and cancer. Computational and experimental biologists are using the prototype infrastructure, thereby aiding the STFC computer scientists in improving the framework and its services.
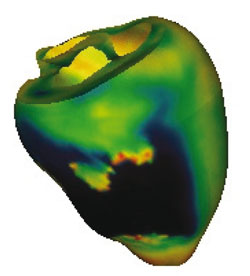
Heart disease and cancer represent the two biggest diseases in the West, and hence have become the focus of intense research within the biomedical community. Computer simulation of whole organs offers the potential to improve our understanding of the causes of these conditions, and eventually to develop new treatment regimes and drugs to reduce their threat to life. The Integrative Biology (IB) project brings together a team uniquely qualified to tackle these problems. It includes researchers from the universities of Oxford, Auckland, the STFC (Science and Technology Facilities Council), Sheffield, Birmingham and Nottingham, University College London, and Leeds. One aspect of this work is the computer simulation of whole organs based on molecular- and cellular-level models, and this requires the application of large-scale computational and data management resources. Another major aspect is the provision of advanced visualization services that allow real-time interactive data analysis on the Grid.
The main components of the Integrative Biology Grid are as follows.
Security access: the Grid Security Infrastructure is employed throughout. The IB project has successfully promoted the Grid computing model and its certificate-based authentication and authorization, not only to computational scientists but also to biologists whose experience of complex computing paradigms is minimal.
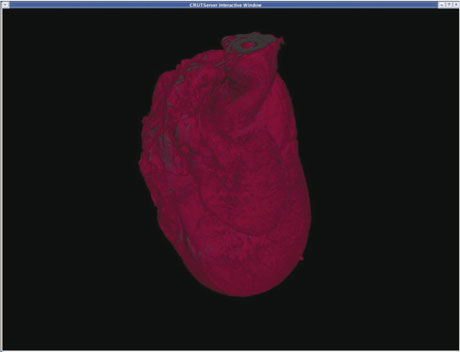
Data management: the use of the Storage Resource Broker for data storage and the use of metadata catalogues for browsing and retrieval have been built into the infrastructure. The complexities are hidden behind user-friendly interfaces within familiar tools.
Interactive access: fundamental to successful analysis and the extension of existing models is the scientist's ability to interactively control, or 'steer', simulations as they are executing. Such experimentation increases understanding the correlation between the parameters under investigation and provides an opportunity for collaboration with colleagues and peers, promoting both the sharing of knowledge and an understanding of the models and the results being produced. A useful dialogue can be established between those working on systems of different scales - from single cells to multi-cell and whole organ models so that a holistic understanding of the chemical, biological and functional processes of diseases can be achieved. The IB interactive services include an elaborate image-based computational steering infrastructure built using the gViz steering library. The steering parameters are bound to appropriate image icons that are placed within the graphics window. A pull-architecture is used to interact with the remote simulation, supporting in situ decisions that are guided by a visual inspection of intermediate data. This image-based steering is developed as a generic portable library usable with OpenGL-based desktop tools. The communication protocols are standards-based and portable.
Visualization: advanced visualization services are built to handle extremely large datasets close to where the data is stored or generated. These include building 3D images from 2D MRI and histology data, 3D isosurfaces, server-side animation generation and viewpoint-dependent animations. Tailor-made generation and rendering services can render up to 0.5GB of data in high resolution and in close to real-time. The results can be sent to the user's desktop using public-domain open-software stacks.
User interfaces: desktop application toolkits familiar to scientists are used to realize interfaces to job submission, workflow, data management, visualization and analysis utilities, and are offered as an integrated data analysis framework. Scientists see an incremental addition in functionality to the tools that they have found effective for their data analysis. This adoption is an essential factor in encouraging widespread take-up of the emerging Integrative Biology Grid.
Virtual Research Environment: the IB services are also deployed within a Virtual Research Environment (VRE) built using portal technologies in the sister project IBVRE. The collaborative research tools built as services within that project will be exported into the main IB project through the generic IB interface.
Links:
http://www.integrativebiology.ox.ac.uk/
http://www.vre.ox.ac.uk/ibvre/
Please contact:
Lakshmi Sastry
Science and Technology Facilities Council, e-Science Centre, UK
Tel: +44 1235 446892
E-mail: m.sastry![]() rl.ac.uk
rl.ac.uk










