by Jane Kernan and Heather J. Ruskin
Large public Microarray databases which store extensive data on different genes, are still in high demand. Integration of the different data types generated poses many problems, not least in terms of quality assessment, scope and interpretation. Old and new paradigms for management co-exist within the field, including Data Warehouses, Database Clusters, Federations and Cloud computing, which need to be sensitive to both historical provision and future directions. So, what is involved?
Microarrays and other high throughput technologies provide the means to measure and analyse expression levels of a large number of genes and samples. In early stage development of microarray technologies, limitations in the initial structure of the datasets rapidly became evident, together with the demand for flexible computational tools and visualisation capability to extract meaningful information. Requirements for shared data access were also recognised. These initial demands have led to new developments in data storage, analysis and visualisation tools, and have facilitated identification of previously undetected patterns in complex datasets. Solving one set of problems is rarely an end in itself, however, and so it has proved in this case, with new data types (typically from Next Generating sequencing methods), now on the increase, the problem is one of reconciling/integrating the different types to maximum advantage.
In any consideration of data quality in the current context, recognition of the major contribution to the success and wide use of microarray technology is due to MIAME, (Minimum Information About a Microarray Experiment), a set of standards for microarray experiment annotation, launched in 1999 by the Microarray Gene Expression Data (MGED) group, (now known as Functional Genomics Data Society (FGED) ). The MGED Ontology working group are continuing to develop a common set of terminologies, the microarray ontology (MO) and Gene Ontology (GO) to facilitate automated querying and exchange of microarray data. Furthermore, the Microarray Quality Control (MAQC) Consortium have developed quality control standards that operate to ensure the reliability of experimental datasets.
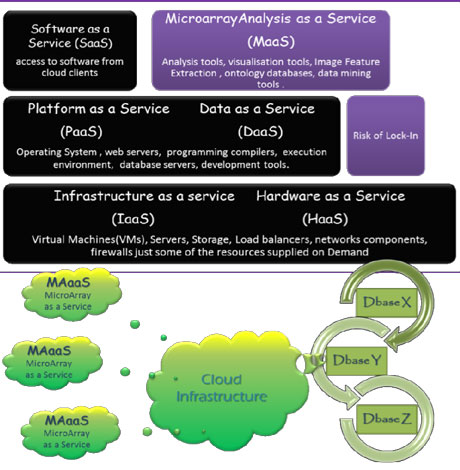
Figure 1 Possible Architecture for Microarray Cloud Computing
Microarray technology has also evolved from the flat file systems and spreadsheets of the early 1990’s to the various database management systems, both public and private, that are accessible through networks and the Internet, sharing data in a more structured and reliable format. With such a variety of web resources, databases, data models, and interfaces, information gathering on properties for a specified gene list is non-trivial as integration is needed to access data, located at numerous remote sources, using different interfaces and query results, which are displayed in various formats. The ideal, clearly, is integration of multiple information sources and a simplified interface, but there is no single solution, to date, which allows comprehensive data on a specific gene to be collated. One federated system, overseen by the National Cancer Institute Center for Biomedical Informatics and Information Technology (NCI-CBIIT) is CaBIG (the Cancer Biomedical Informatics Grid http://cabig.nci.nih.gov/), where a suite of tools provides for simplification of data input and sharing across the Cancer Biomedical Informatics Grid, using a federated model of local installations. A Cloud hosting environment for health care data, (attracting a monthly subscription), is also provided.
Is cloud computing the answer to integration? Does cloud computing make economic sense? Its selling point is that using a cloud computing facility means you do not have to pay for distribution, infrastructure installation or maintenance costs. You only pay for what you use. Software updates and compatibilities are no longer an issue and this is achieved by use of Virtual Machines (VMs) . VMs provide the ability for resource pooling. One cloud computing service provider is Amazon (Elastic Cloud Computing EC2) offers a variety of bioinformatics-oriented virtual machine images as well as providing several large genomic datasets in its cloud (copies of GeneBank DB and Genome databases from Ensembl), once portability and interoperability standards are introduced this may be a viable option.
When choosing to migrate to a cloud one of the main concerns is that important data will reside on a third parties server, other concerns are, security and lack of control over the system, the trust-worthiness of the service provider, and the lack of portability standards, which may heighten the risk of company lock-in for customers wishing to move vendors. How simple a process is moving vendors going to be? Governing authorities such as the IEEE Standards Association (IEEE-SA) have formed two new Working Groups (WGs) IEEE P2301 (Cloud Portability and Interoperability) and IEEE P2302 (Standard for Inter cloud Interoperability and Federation) to try and standardise the approaches of critical areas such as portability, interoperability interfaces and file formats.
Migrating to cloud computing is not a trivial task, so the question is can Cloud with its combination of Computational Power, Big Data and Distributed Storage, Horizontal Scalability and Reliability be harnessed to provide a one-stop-shop for Microarray Analysis? Can complex multi-faceted searches on a distribution of connected databases be supplied on demand to microarray researchers through a Microarray Analysis-as-a-Service type client process (which is a "one-to-many" model whereby an application is shared across multiple clients). This research is on-going and future work will report on interoperability and migration issues of these biological databases when combined with the latest cloud technologies.
Links:
http://opencloudmanifesto.org/Cloud_Computing_Use_Cases_Whitepaper-4_0.pdf http://www.biomedcentral.com/1471-2105/12/356
Please contact:
Jane Kernan
CloudCORE Research Centre
Dublin City University, Ireland
E-mail:










