by Manolis Tsiknakis
ACGT (Advancing Clinico-Genomic Trials on cancer) is an FP6 integrated project focusing on the development of a semantic Grid infrastructure to support multicentric, post-genomic clinical trials. This will enable discoveries in the laboratory to be quickly transferred to clinical management and the treatment of patients.
Recent advances in research methods and technology have resulted in an explosion of information and knowledge about cancers and their treatment. Exciting new research on the molecular mechanisms that control cell growth and differentiation has resulted in a significant improvement in our understanding of the fundamental nature of cancer cells, and has suggested valuable new approaches to cancer diagnosis and treatment. Despite these advances, the lack of a common infrastructure has prevented clinical research institutions from mining and analysing disparate data sources. This inability to share technology and data developed by different organizations is severely hampering the research process.
Post-Genomic Clinical Trials
The ACGT project has been structured to deal with this problem. The ERCIM office is the administrative project coordinator, while FORTH is responsible for scientific coordination. The project has selected two cancer domains (breast cancer and Wilm's tumour or pediatric nephroblastoma) and has defined specific trials which are feeding the requirement analysis and elicitation phase of the project. A third trial is also included, which focuses on the reuse of multilevel biomedical data produced in the previous two trials and the integration of advanced technology (including interactive visualization, virtual reality technology and in silico tumour growth simulations). Here the objective is to explore simulated predictions of tumour growth and treatment response.
The Breast Cancer Trial
Breast cancer is both genetically and histopathologically heterogeneous, and the mechanisms underlying its development remain largely unknown. The ACGT Test of Principle (TOP) study aims to identify biological markers associated with pathological complete response to anthracycline therapy (epirubicin), one of the most active drugs used in breast cancer treatment. Supported by in vitro and preliminary in vivo data, this study is designed to test prospectively the value of topo II alpha gene amplification and protein overexpression in predicting the efficacy of anthracyclines.
These clinical trials are multicentric (many different research organizations are participating), and post-genomic, meaning they require generation, management, integrated access, processing and analysis of multilevel biomedical data, including transcriptomic, proteomic and imaging data. As a result, the ultimate objective of the ACGT project is the development of semantic Grid infrastructure that offers high-level tools and techniques for the distributed mining and extraction of knowledge from data repositories available on the Grid. This infrastructure will make use of semantic descriptions of components and data and will offer knowledge discovery services in the domain of cancer research. Special emphasis is given to the trust that needs to be embedded in the platform, and to relevant ethical issues, thus creating optimal conditions for service uptake.
Since we see the requirements engineering process as a structured set of activities that will lead to the fulfillment of the final system requirements, an iterative method has been adopted, based mainly on scenarios and prototyping. Explicit scenarios have been developed that represent documented user needs and also provide a technology-driven description of the requirements of the system under design, as understood by experienced technological experts.
Initial System Architecture
From a detailed analysis of documented user requirements, it is apparent that a complex technical infrastructure must be developed if support for integrated access, analysis and visualization of multilevel, heterogeneous data is to be provided. A detailed analysis of the scientific and functional requirements of the ACGT infrastructure was performed, together with an analysis of the current state of the art in terms of technological infrastructure, data resources, data representation, exchange standards and ontologies.
With respect to the state of the art, the myGrid project (http://www.mygrid. org.uk) is focusing on providing in silico support for experimental research, while the cancer Biomedical Informatics Grid (caBIG - https://cabig.nci.nih. gov/) is creating a virtual community within which resources can be shared and the key issues of cyber infrastructure tackled.
From a technical point of view, the requirements identified can be met using a federated, multilayer, service-oriented and ontology-driven architecture. The ACGT project decided to build on open software frameworks based on the WS-Resource Framework (WSRF) and Open Grid Service Architecture (OGSA), which are the de facto standards in Grid computing. These standards are implemented in the selected middleware, namely Globus Toolkit 4 (GT4) (http://www.globus.org) and Gridge (http://fury.man.poznan.pl/gridge/).
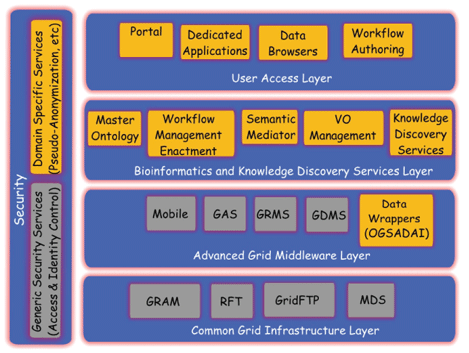
An overview of the ACGT system layered architecture is given in the figure, and includes the following layers:
-
Common Grid Infrastructure Layer: this comprises the basic "Grid engine" for accessing remote resources in a Grid environment. It provides a common interface for Grid resources used by higher-level services
-
Advanced Grid Middleware Layer: this comprises advanced Grid services, which operate on sets of lower-level services to provide more advanced functionality
-
Bioinformatics and Knowledge Discovery Service Layer: this includes all the ACGT-specific services, such as the ACGT Master Ontology, the Clinical Trial on Cancer Metadata Services, semantic mediation services and distributed and privacy-preserving data-mining and knowledge discovery services
-
User Access Layer: this allows users to realize complex biomedical applications by combining basic services from the underlying layers and exploiting the resources and data provided by the research centres that form different CT Virtual Organizations (VOs)
-
Security Layer: access rights, security and trust-building are issues addressed by this layer.
Biomedical Grid Intelligence
In a "Grid-enabled" data-sharing VO, datasets may not be well known amongst all VO participants. To integrate highly fragmented and isolated data sources, we need semantics in order to answer higher-level questions. It therefore becomes critically important to describe the context in which the data was captured. We describe this contextualization of the data as metadata. Semantic integration in ACGT thus relies on metadata publishing and ontologies.
We see as our main future research challenge in ACGT the development of an infrastructure that is able to produce, use and deploy knowledge as a basic element of advanced applications. This will mainly constitute a Biomedical Knowledge Grid. Metadata is critical to achieving such an objective. We use OWL-S to develop metadata and service ontologies for describing Grid Services so that they might be discovered, explained, composed and executed automatically.
Our initial investigations have also revealed the need for a sophisticated model of provenance, since the use of both elementary and advanced workflows (workflows containing other workflows), is becoming a very important goal in our R&D work. This requirement also involves maintaining complex metadata relating to workflows in the ACGT Grid middleware.
Link:
http://www.eu-acgt.org
Please contact:
Manolis Tsiknakis
ICS-FORTH, Greece
Tel: +30 2810 391690
E-mail: tsiknaki![]() ics.forth.gr
ics.forth.gr










