by Leonardo Candela and Pasquale Pagano
Data sharing has been an emerging topic since the 1980’s. Science evolution – e.g. data-intensive, open science, science 2.0 – is revamping this discussion and calling for data infrastructures capable of properly managing data sharing and promoting extensive reuse. ‘gCube’, a software system that promotes the development of data infrastructures, boasts the distinguishing feature of providing its users with Virtual Research Environments where data sharing and reuse actually happens.
gCube - a software system designed to enable the creation and operation of an innovative typology of data infrastructure - leverages Grid, Cloud, digital library and service-orientation principles and approaches to deliver data management facilities as-a-service. One of its distinguishing features is that it can serve the needs of diverse communities of practice by providing each with one or more dedicated, flexible, ready-to-use, web-based working environments, i.e. Virtual Research Environ-ments [1].
gCube provides its users with services for seamless access to species data, geospatial data, statistical data and semi-structured data from diverse data providers and information systems. These services can be exploited both via web-based graphical user interfaces and web-based protocols for programmatic access, e.g., OAI-PMH, CSW, SDMX.
For species data, gCube is equipped with a Species Data Discovery (SDD) Service [2] which mediates over a number of data sources including taxonomic information, checklists and occurrence data. The service is equipped with plug-ins interfacing with major information systems such as Catalogue of Life, Global Biodiversity Information Facility, Integrated Taxonomic Information System, Interim Register of Marine and Nonmarine Genera, Ocean Biogeographic Information System, World Register of Marine Species. To expand the number of information systems and data sources integrated into SDD, the VRE data manager can simply implement (or reuse) a plug-in. Each plug-in can interact with an information system or database by relying on a standard protocol, e.g., TAPIR, or by interfacing with its proprietary protocol. Plug-ins mediate queries and results from the language and model envisaged by SDD to the requirements of a particular database. SDD promotes a data discovery mechanism based on queries containing either the scientific name or common name of a species. Furthermore, to tackle issues arising from inconsistency in taxonomy among data sources, the service supports an automatic query expansion mechanism, i.e. the query could be augmented with ‘similar’ species names. Discovered data is presented in a homogenized form, e.g., in a typical Darwin Core format.
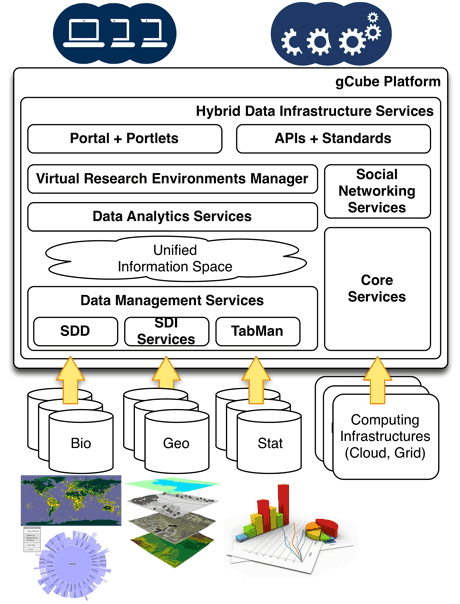
Figure 1: The gCube System Architecture.
For geospatial data, gCube is equipped with services generating a Spatial Data Infrastructure compliant with OGC standards. In particular, it offers a catalogue service enabling the seamless discovery of and access to every geospatial resource registered or produced via gCube services. These resources include physical and biochemical environmental parameters, such as temperature and chlorophyll, species distribution and occurrence maps, and other interactive maps. Some of these resources are obtained by interfacing with existing Information Systems including FAO GeoNetwork, myOcean and World Ocean Atlas. New resources can be added by linking data sources to the SDI via standards or ad-hoc mediators. On top of the resulting information space, gCube offers an environment for identifying resources and overlays them through an innovative map container that caters for sorting, filtering, and data inspection further to standard facilities such as zoom in.
For statistical data, the infrastructure is equipped with a dedicated statistical environment supporting the whole lifecycle of statistical data management, including data ingestion, curation, analysis and publication. This environment provides its users with facilities for creating new datasets and code lists by using sources like CSV or an SDMX repository, curating the datasets (by using controlled vocabularies and code lists, defining data types and correcting errors), manipulating the datasets with standard operations like filtering, grouping, and aggregations, analysing the datasets with advanced mining techniques, such as trend and outlier detection, producing graphs from the datasets, and finally publishing datasets in an SDMX registry for future use.
On top of the unified information space which is underpinned by the facilities described above, gCube provides its users with social networking facilities [2] and data analytics facilities [3].
Link:
http://www.gcube-system.org
References:
[1] L. Candela, D. Castelli, P. Pagano (2013) Virtual Research Environments: an Overview and a Research Agenda. Data Science Journal, 12:GRDI75–GRDI81, 2013.
[2] M. Assante et al. (2014) A Social Networking Research Environment for Scientific Data Sharing: The D4Science Offering. The Grey Journal, Vol. 10, Number 2, 2014
[3] L. Candela et al. (2014) An infrastructure-oriented approach for supporting biodiversity research. Ecological Informatics, DOI: 10.1016/j.ecoinf.2014.07.006, Elsevier
Please contact:
Pasquale Pagano
ISTI-CNR, Italy
E-mail:











