by Peter Murray-Rust (University of Cambridge)
Scholarly publications, especially science and medicine, have huge amounts of untapped knowledge, but it’s a technical challenge to extract it and there’s a political fight in Europe as to whether we can legally do it.
About three million peer-reviewed scholarly publications and technical reports, especially in life science and medicine, are published each year – one every 10 seconds. Many are filled with facts (species, diseases, drugs, countries, organisations) resulting from about one trillion USD of funded research. But they aren’t properly used – for example the Zika outbreak was predicted 30 years ago [1] but in a scanned PDF behind a paywall so was never broadcast. Computers are essential to process this data – but there are major problems: the complexity of semi-structured information and socio-political conflict is being played out in Brussels even as I write this [2].
Scientists write in narrative text, with embedded data and images and no thought for computer processing. Most text is born digital (Word, TeX) and straightforward to process but turned into PDF. Data is collected from digital instruments, summarised to diagrams and turned into pixels (PNG, JPG) with total loss of data – even from summary diagrams (plots). We know of researchers who spend their whole time turning this back into computable information.
However, it’s still possible to recover a vast amount of data with heuristics such as natural language processing and diagram processing. With Shuttleworth Foundation funding I’ve created ContentMine.org [3] to read the whole scientific literature and extract the facts on a daily basis. We’ve spent two years creating the code and pipeline and are now starting to process open and subscription articles automatically – and this can extend to theses and reports.
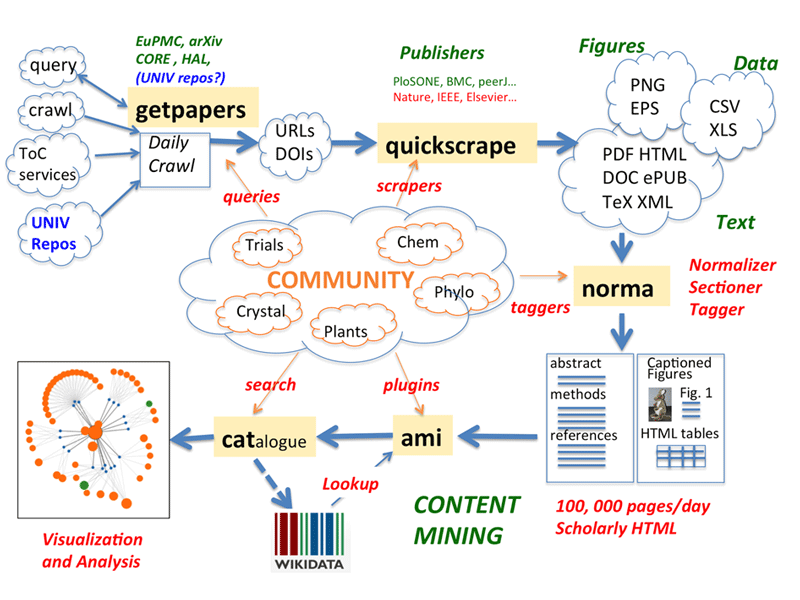
Figure 1: The ContentMine pipeline. Articles can be ingested by systematic crawling, push from a TableOfContents service, or user-initiaited query (getpapers and/or quickscrape). They are normalised to XHTML, annotated by section and facts extracted by ‘plugins’ and sent to public repositories (Wikidata, Zenodo). All components are Open Source/data.
The pipeline covers many tasks including crawling and scraping websites, using APIs or papers already aggregated. Papers often need normalising from raw HTML to structured, annotated XHTML and marking up the sections (‘Introduction’, ‘Methods’, ‘Results’, etc ) is an important way of reducing false positives. Captions for tables and figures are often the most important parts of some articles. We then search the text by discipline-specific plugins, most commonly using simple dictionaries enhanced by Wikidata. These often exist in current disciplines – e.g., ICD-10 for disease – and increasingly we can extract them directly from Wikidata. More complex tools are required for species and chemistry. And we have pioneered automatic methods for interpreting images of phylogenetic trees and constructed a supertree for 4,500 articles.
Among the sources are EuropePubMedCentral – over one million open articles on life science and medicine, converted into XML. Our getpapers tool directly uses EPMC’s search API and feeds text for conversion to scholarlyHTML. We can also get metadata from Crossref and scrape sites directly with per-publisher scrapers – it takes less than an hour to create a new one.
We see Wikidata as the future of much scientific fact, and cooperate with them by creating enhanced dictionaries for searching and also providing possible new entries. The Wikidata-enhanced facts will be stored in the public Zenodo database for all to use. Since facts are uncopyrightable we expect to extract over 100 million per year.
Text and data mining (or ContentMining) has been seen as a massive public good[5]. Sir Mark Walport, director of the Wellcome Trust, said "This is a complete no-brainer. This is scholarly research funded from the public purse, largely from taxpayer and philanthropic organisations. The taxpayer has the right to have maximum benefit extracted and that will only happen if there is maximum access to it."
But there’s huge politico-legal opposition, because the papers are copyrighted, normally by the publishers who see mining as a new revenue stream, even though they have not developed the technology. Innovative scientists carrying out mining risk their universities being cut off by publishers. The UK has pioneered reform to allow mining for non-commercial research, but it was strongly opposed by publishers and there’s little effective practical support. In 2013 organisations such as national libraries, funders, and academics were opposed by rightsholders (‘Licences for Europe’) leading to an impasse. The European parliament has tried to reform copyright, but recommendations have been heavily watered down by the commission and leaks suggest that formalising the market for exploitation by publishers will be emphasised at the expense of innovation and freedom.
We desperately need open resources – content, dictionaries, software, infrastructure. The UK has led but not done enough. France is actively deciding on its future. Within two years decisions will become effectively irrevocable. Europe must choose whether it wants mining to be done by anyone, or controlled by corporations.
Links:
[L1] http://www.nytimes.com/2015/04/08/opinion/yes-we-were-warned-about-ebola.html?_r=0
[L2] http://kluwercopyrightblog.com/2016/07/20/julia-reda-mep-discusses-harmonisation-copyright-law-ip-enforcement-brexit/
[L3] http://contentmine.org
[L4] http://www.statewatch.org/news/2016/aug/eu-com-copyright-draft.pdf
[L5] https://www.jisc.ac.uk/news/text-mining-promises-huge-economic-and-research-benefit-but-barriers-limit-its-use-14-mar-2012
Reference:
P Murray-Rust, J Molloy, and D Cabell: “Open content mining”, in Proc. of The First OpenForum Academy Conference, pp. 57-64. OpenForum Europe LTD, 2012.
Please contact:
Peter Murray-Rust, University of Cambridge, UK
+44 1223 336432,











