by Gunnar Klau
CWI researchers are developing the Heinz family of algorithms to explore big life sciences data in the context of biological networks. Their methods recently pointed to a novel hypothesis about how viruses hijack signalling pathways.
High-throughput technologies are generating more and more measurements of cellular processes in health and disease, leading to ever-increasing amounts of “omics” data. The main challenge is now to interpret and understand these massive data sets in order to ultimately answer important biological and biomedical questions. This is a complex challenge, however, because important signals are hidden by high noise levels and heterogeneity of samples and diseases. Inspecting big life sciences data in the context of biological networks helps to address this challenge. To this end, researchers at CWI in Amsterdam are developing the Heinz family of algorithms to analyse and explore genome-scale measurements within biological networks.
The Heinz family of algorithms
The Heinz project started in 2007 as a collaboration with University of Würzburg’s Biocenter. The first algorithmic prototype, Heinz 1.0 [1], was presented in 2008 at ISMB, the premier conference on computational biology. There, the work received the outstanding paper award for achieving a breakthrough in computing optimal subnetwork modules by introducing and exploiting a relation to graph theory. Since then, the method has been improved and several variations have been presented:
- Heinz, the workhorse method, is currently at version 2.0 [L1]. It takes as input a set of gene scores and finds an optimal active subnetwork module with respect to these scores. This is a connected subnetwork where the sum of the gene scores is maximal. Finding such a module is an NP-hard problem. Heinz computes provably optimal modules using advanced techniques from mathematical optimization. While Heinz 1.0 exploited the close relation of the underlying Maximum-Weight Connected Subgraph (MWCS) problem to the Prize-Collecting Steiner Tree (PCST) problem and relied on PCST codes, Heinz 2.0 directly solves MWCS using a recursive graph-decomposition scheme into bi- and tri-connected components and a dedicated branch-and-cut algorithm.
- BioNet [L2] is an R package that provides an easy-to-use interface to Heinz. Provided with raw data, e.g., from RNA-Seq measurements, it generates the input score files needed by Heinz. The scores are based on a statistically sound decomposition of p-values describing the measurements into signal and noise components.
- Heinz has been adapted to answer a variety of research questions involving the interpretation of differential gene expression, GWAS, metabolomics, proteomics and metagenomics data.
- xHeinz [L3] is a recent addition that computes conserved cross-species modules. In a cooperation with the Netherlands Cancer Institute (NKI), xHeinz was used to provide evidence that the differentiation process of the recently discovered Th17 cell type, which plays an important role for the immune system, is conserved between mouse and human [2].
- eXamine, a visual analytics app for exploring Heinz results in the popular Cytoscape visualization platform, was developed in a cooperation with Eindhoven University of Technology. The tool makes it easy to explore annotated Heinz modules, for example, in the context of Gene Ontology or pathway enrichment.
All software in the Heinz family is open source.
Case study on virally deregulated s ignalling
The Human Cytomegalovirus (HCMV) is a specific type of herpes virus with a high prevalence of 60% among humans. The interplay of HCMV infections with many diseases, including cancer, is an important topic of biomedical research. In a collaboration within the Amsterdam Institute for Molecules, Medicines and Systems, Heinz and eXamine were used to study a module that is activated by an HCMV-encoded G-protein coupled receptor. See Figure 1 for an illustration of the optimal Heinz module along with enriched functional and pathway categories using eXamine. Using the tools from the Heinz family, the researchers have been able to formulate a new hypothesis about deregulated signalling of β-catenin by viral receptor proteins. Parts of this new hypothesis have now been verified experimentally and have led to targeted follow-up studies, which are currently under way.
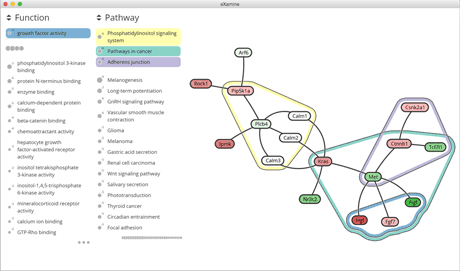
Figure 1: Optimal Heinz module along with enriched functional and pathway categories using eXamine.
Future
Current research includes the application to cancer genomics data. Here, the task is to extract subnetworks that show exclusive mutation patterns in the samples. A long term research goal is to move towards more dynamic descriptions of cellular mechanisms.
Links:
[L1] http://software.cwi.nl/heinz
[L2] http://bionet.bioapps.biozentrum. uni-wuerzburg.de
[L3] http://software.cwi.nl/xheinz
References:
[1] M. Dittrich et al.: “Identifying functional modules in protein-protein interaction networks: an integrated exact approach”, Bioinformatics 24(13):i223-i231, 2008.
[2] M. El-Kebir et al.: “xHeinz: an algorithm for mining cross-species network modules under a flexible conservation model”, Bioinformatics 31(19):3147-55, 2015.
[3] K. Dinkla et al.: “eXamine: Exploring annotated modules in networks”, BMC Bioinformatics 15:201, 2014.
Please contact:
Gunnar W. Klau
CWI, currently Fulbright Visiting Professor at Brown University
Tel: +31 20 592 4012
E-mail:











