by Alexander Schönhuth and Tobias Marschall
Detecting genetic variants is like spotting tiny sequential differences among gigantic amounts of text fragment data. This explains why some variants are extremely hard to detect or have even formed blind spots of discovery. At CWI, we have worked on developing new tools to eliminate some of these blind spots. As a result, many previously undiscoverable genetic variants now form part of an exhaustive variant catalogue based on the Genome of the Netherlands project data.
In 2007, the advent of "next-generation sequencing" technologies revolutionized the field of genomics. It finally became affordable to analyse large numbers of individual genomes, by breaking the corresponding DNA into fragments and sequencing those fragments, yielding “sequencing reads”. All of this is now happening at surprisingly – nearly outrageously – low cost and high speed. Advances in terms of cost and speed, paired with the relatively short length of the fragments (in comparison to “first-generation sequencing”) comes at a price, however. First, the rapid pile-up of sequencing reads makes for a genuine “big data” problem. Second, the reduced fragment length yields even more complex scientific riddles than in “first-generation sequencing” times. Overall, the resulting computational problems are now harder both from theoretical and practical points of view. Despite – or possibly owing to – the incredible mass of data, certain genetic variants stubbornly resist detection and form blind spots of genetic variant discovery due to experimental and statistical limitations. Note that, in the absence of adequate methods to detect them, the first question to ask is: do these variants even exist in nature?
The presence of possible blind spots has not kept researchers from analysing these gigantic haystacks of sequence fragments. A prominent example of such an effort is the "Genome of the Netherlands" project [2], which has aimed at providing an exhaustive summary of genetic variation for a consistent population. Launched in 2010, it is both one of the earliest population-scale sequencing projects, and still one of the largest of its kind -- overall, the fragment data amounts to about 60 terabytes. The analysis of sequencing data is further enhanced by sequencing related individuals – either family trios or (twin) quartets – which allows the researchers to study transmission of variants and variant formation within one generation [3]. The resulting catalogue of variants establishes an invaluable resource, not only for the Dutch, but also for closely related European populations regarding association of disease risks with DNA sequence variation, and personalized medicine in general.
At CWI, as members of the Genome of the Netherlands project, we have succeeded in eliminating a prominent discovery blind spot, thereby contributing large numbers of previously undiscoverable genetic variants. We achieved this by reversing a common variant discovery workflow – usually, large amounts of seemingly ordinary looking sequence fragments are removed, which turns a big into a small data problem and renders fragment analysis a lot easier. In contrast, we process all data [1]: in other words, instead of removing large amounts of hay and, with it, considerable amounts of needles that are too tiny to be easily spotted, we re-arrange the entire haystack such that even the tiny needles stick out. We have developed a "statistical magnet" that pulls the tiny needles to the surface.
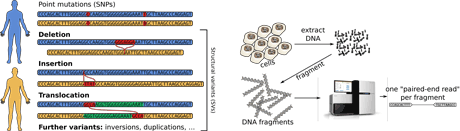
Figure 1: Left: Different classes of genetic variants in human genomes. Right: Next-generation sequencing, only after breaking up DNA in small fragments, one can read the DNA – however, deletions and insertions of length 30-200 letters now are very difficult to spot. We have eliminated this blind spot in discovery by developing new algorithms.
The key to success has been the development of an ultra-fast algorithm that empowers the application of this magnet even on such massive amounts of sequence fragments. In summary, the combination of a sound statistical machinery with a highly engineered algorithm allows for implementation of a reversed discovery workflow.
As a result, the Genome of the Netherlands project is the first of its kind to exhaustively report on the corresponding class of genetic variants, previously termed “twilight zone deletions and insertions”, but which now enjoy somewhat more daylight.
In future work, we are also planning to eliminate this blind spot in somatic variant discovery, which will likely reveal large amounts of so far undetected cancer-causing genetic variants, and will hopefully shed considerable light on cancer biology as well.
Links:
http://homepages.cwi.nl/~as
http://www.nlgenome.nl
References:
[1] T. Marschall, I. Hajirasouliha, A. Schönhuth: “MATE-CLEVER: Mendelian-inheritance-aware discovery and genotyping of midsize and long indels”, Bioinformatics 29(24):3143-3150, 2013.
[2 ] The Genome of the Netherlands Consortium: “Whole-genome sequence variation, population structure and demographic history of the Dutch population”, Nature Genetics 46(8):818-825, 2014.
[3] W. Kloosterman, et al.: “Characteristics of de novo structural changes in the human genome”, Genome Research 25:792-801, 2015.
Please contact:
Alexander Schönhuth
CWI, The Netherlands
E-mail:
Tobias Marschall was a postdoc at CWI from 2011-2014. Since 2014, he holds an appointment as assistant professor at the Center for Bioinformatics at Saarland University and the Max Planck Institute for Informatics in Saarbrücken,
Germany











